
Quick start
1.
Download
git clone https://github.com/Ensembl/ensembl-vep.git
2. Install
cd ensembl-vep
perl INSTALL.pl
3.
Test
./vep -i examples/homo_sapiens_GRCh38.vcf --cache
Download documentation in PDF format
Tutorial
Download and install
Download
What's new in release 112
Installation
Using VEP in Windows
Docker
Data formats
Running VEP
Options
Annotation sources
Caches
GFF/GTF files
FASTA files
Databases
Filtering results
Variant Effect Predictor Command line
VEP
Use VEP to analyse your variation data locally. No limits, powerful, fast and extendable,
command line VEP is the way to get the most out of VEP and Ensembl.
VEP is a powerful and highly configurable tool - have a browse through the documentation.
You might also like to read up on the data formats that VEP uses, and the different ways
you can access genome data. The VEP script can annotate your variants with custom data,
be extended with plugins, and use powerful filtering to find biologically interesting results.
Beginners should have a run through the tutorial, or try the web interface first.
If you use VEP in your work, please cite our latest publication McLaren et. al. 2016
(doi:10.1186/s13059-016-0974-4 )
Any questions? Send an email to the Ensembl developers' mailing list or contact the
Ensembl Helpdesk.
Documentation
contents

Input
Output
Running filter_vep
Writing filters
Custom annotations
Data formats
Options
Plugins
Existing plugins
Using plugins
Examples & use cases
Example commands
gnomAD exomes and genomes
Citations and VEP users
Other information
Performance
Multiple assemblies
Summarising annotation
HGVS notations
RefSeq transcripts
FAQ
General questions
Web VEP questions
Command line VEP questions
!

Variant Effect Predictor Tutorial
Install VEP
Have you downloaded VEP yet? Use git to clone it:
git clone https://github.com/Ensembl/ensembl-vep
cd ensembl-vep
VEP uses "cache files" or a remote database to read genomic data. Using cache files gives
the best performance - let's set one up using the installer:
perl INSTALL.pl
Hello! This installer is configured to install v112 of the
Ensembl API for use by VEP.
It will not affect any existing installations of the Ensembl API
that you may have.
It will also download and install cache files from Ensembl's FTP
server.
Checking for installed versions of the Ensembl API...done
It looks like you already have v112 of the API installed.
You shouldn't need to install the API
Skip to the next step (n) to install cache files
Do you want to continue installing the API (y/n)?
If you haven't yet installed the API, type "y" followed by enter, otherwise type "n" (perhaps if
you ran the installer before). At the next prompt, type "y" to install cache files
Do you want to continue installing the API (y/n)? n
- skipping API installation
VEP can either connect to remote or local databases, or use local
cache files.
Cache files will be stored in /nfs/users/nfs_w/wm2/.vep
Do you want to install any cache files (y/n)? y
Downloading list of available cache files
The following species/files are available; which do you want (can
specify multiple separated by spaces):
1 : ailuropoda_melanoleuca_vep_112_ailMel1.tar.gz
2 : anas_platyrhynchos_vep_112_BGI_duck_1.0.tar.gz
3 : anolis_carolinensis_vep_112_AnoCar2.0.tar.gz
...
42 : homo_sapiens_vep_112_GRCh38.tar.gz
...
?
Type "42" (or the relevant number for homo_sapiens and GRCh38) to install the cache for
the latest human assembly. This will take a little while to download and unpack! By default
VEP assumes you are working in human; it's easy to switch to any other species using --
species [species].
? 42
- downloading
https://ftp.ensembl.org/pub/release-112/variation/vep/homo_sapien

s_vep_112_GRCh38.tar.gz
- unpacking homo_sapiens_vep_112_GRCh38.tar.gz
Success
By default VEP installs cache files in a folder in your home area ($HOME/.vep); you can
easily change this using the -d flag when running the installer. See the installer
documentation for more details.
Run VEP
VEP needs some input containing variant positions to run. In their most basic form, this
should just be a chromosomal location and a pair of alleles (reference and alternate). VEP
can also use common formats such as VCF and HGVS as input. Have a look at the Data
formats page for more information.
We can now use our cache file to run VEP on the supplied example file
examples/homo_sapiens_GRCh38.vcf, which is a VCF file containing variants from the
1000 Genomes Project, remapped to GRCh38:
./vep -i examples/homo_sapiens_GRCh38.vcf --cache
2013-07-31 09:17:54 - Read existing cache info
2013-07-31 09:17:54 - Starting...
ERROR: Output file variant_effect_output.txt already exists.
Specify a different output file
with --output_file or overwrite existing file with --
force_overwrite
You may see this error message if you've already run VEP in the same directory. VEP tries
not to trample over your existing files unless you tell it to. So let's tell it to using --
force_overwrite
./vep -i examples/homo_sapiens_GRCh38.vcf --cache --
force_overwrite
By default VEP writes to a file named "variant_effect_output.txt" - you can change this file
name using -o. Let's have a look at the output.
head variant_effect_output.txt
## ENSEMBL VARIANT EFFECT PREDICTOR v112.0
## Output produced at 2017-03-21 14:51:27
## Connected to homo_sapiens_core_112_38 on ensembldb.ensembl.org
## Using cache in /homes/user/.vep/homo_sapiens/112_GRCh38
## Using API version 112, DB version 112
## polyphen version 2.2.2
## sift version sift5.2.2
## COSMIC version 78
## ESP version 20141103
## gencode version GENCODE 25
## genebuild version 2014-07
## HGMD-PUBLIC version 20162
## regbuild version 16
## assembly version GRCh38.p7
## ClinVar version 201610
## dbSNP version 147
## Column descriptions:
## Uploaded_variation : Identifier of uploaded variant
## Location : Location of variant in standard coordinate format
(chr:start or chr:start-end)
## Allele : The variant allele used to calculate the consequence

## Gene : Stable ID of affected gene
## Feature : Stable ID of feature
## Feature_type : Type of feature - Transcript, RegulatoryFeature
or MotifFeature
## Consequence : Consequence type
## cDNA_position : Relative position of base pair in cDNA
sequence
## CDS_position : Relative position of base pair in coding
sequence
## Protein_position : Relative position of amino acid in protein
## Amino_acids : Reference and variant amino acids
## Codons : Reference and variant codon sequence
## Existing_variation : Identifier(s) of co-located known
variants
## Extra column keys:
## IMPACT : Subjective impact classification of consequence type
## DISTANCE : Shortest distance from variant to transcript
## STRAND : Strand of the feature (1/-1)
## FLAGS : Transcript quality flags
#Uploaded_variation Location Allele Gene
Feature Feature_type Consequence ...
rs7289170 22:17181903 G ENSG00000093072
ENST00000262607 Transcript synonymous_variant ...
rs7289170 22:17181903 G ENSG00000093072
ENST00000330232 Transcript synonymous_variant ...
The lines starting with "#" are header or meta information lines. The final one of these
(highlighted in blue above) gives the column names for the data that follows. To see more
information about VEP's output format, see the Data formats page.
We can see two lines of output here, both for the uploaded variant named rs7289170. In
many cases, a variant will fall in more than one transcript. Typically this is where a single
gene has multiple splicing variants. Here our variant has a consequence for the transcripts
ENST00000262607 and ENST00000330232.
In the consequence column, we can see the consequence term synonymous_variant. This
is terms forms part of an ontology for describing the effects of sequence variants on
genomic features, produced by the Sequence Ontology (SO) . See our predicted data
page for a guide to the consequence types that VEP and Ensembl uses.
Let's try something a little more interesting. SIFT is an algorithm for predicting whether a
given change in a protein sequence will be deleterious to the function of that protein. VEP
can give SIFT predictions for most of the missense variants that it predicts. To do this,
simply add --sift b (the b means we want both the prediction and the score):
./vep -i examples/homo_sapiens_GRCh38.vcf --cache --
force_overwrite --sift b
SIFT calls variants either "deleterious" or "tolerated". We can use the VEP's filtering tool to
find only those that SIFT considers deleterious:
./filter_vep -i variant_effect_output.txt -filter "SIFT is
deleterious" | grep -v "##" | head -n5
#Uploaded_variation Location Allele Gene
Feature ... Extra
rs2231495 22:17188416 C ENSG00000093072
ENST00000262607 ... SIFT=deleterious(0.05)
rs2231495 22:17188416 C ENSG00000093072
ENST00000399837 ... SIFT=deleterious(0.05)
rs2231495 22:17188416 C ENSG00000093072
ENST00000399839 ... SIFT=deleterious(0.05)
rs115736959 22:19973143 A ENSG00000099889
ENST00000263207 ... SIFT=deleterious(0.01)
Note that the SIFT score appears in the "Extra" column, as a key/value pair. This column

can contain multiple key/value pairs depending on the options you give to VEP. See the
Data formats page for more information on the fields in the Extra column.
You can also configure how VEP writes its output using the --fields flag.
You'll also see that we have multiple results for the same gene, ENSG00000093072. Let's
say we're only interested in what is considered the canonical transcript for this gene (--
canonical), and that we want to know what the commonly used gene symbol from HGNC is
for this gene (--symbol). We can also use a UNIX pipe to pass the output from VEP directly
into the filtering tool:
./vep -i examples/homo_sapiens_GRCh38.vcf --cache --
force_overwrite --sift b --canonical --symbol --tab --fields
Uploaded_variation,SYMBOL,CANONICAL,SIFT -o STDOUT | \
./filter_vep --filter "CANONICAL is YES and SIFT is deleterious"
...
#Uploaded_variation SYMBOL CANONICAL SIFT
rs2231495 CECR1 YES deleterious(0.05)
rs115736959 ARVCF YES deleterious(0.01)
rs116398106 ARVCF YES deleterious(0)
rs116782322 ARVCF YES deleterious(0)
... ... ... ...
rs115264708 PHF21B YES deleterious(0.03)
So now we can see all of the variants that have a deleterious effect on canonical
transcripts, and the symbol for their genes. Nice!
For species with an Ensembl database of variants, VEP can be configured to annotate your
input with identifiers and frequency data from variants co-located with your input data. For
human, VEP's cache contains frequency data from 1000 Genomes, NHLBI-ESP and ExAC.
Since our input file is from 1000 Genomes, let's add frequency data using --af_1kg:
./vep -i examples/homo_sapiens_GRCh38.vcf --cache --
force_overwrite --af_1kg -o STDOUT | grep -v "##" | head -n2
#Uploaded_variation Location Allele Gene
Feature ... Existing_variation Extra
rs7289170 22:17181903 G ENSG00000093072
ENST00000262607 ... rs7289170
IMPACT=LOW;STRAND=-1;AFR_AF=0.2390;AMR_AF=0.2003;EAS_AF=0.0456;EU
R_AF=0.3211;SAS_AF=0.1401
We can see frequency data for the AFR, AMR, EAS, EUR and SAS continental population
groupings; these represent the frequency of the alternate (ALT) allele from our input (G in
the case of rs7289170). Note that the Existing_variation column is populated by the
identifier of the variant found in the VEP cache (and that it corresponds to the identifier from
our input in Uploaded_variation). To retrieve only this information and not the frequency
data, we could have used --check_existing (--af_1kg silently switches on --check_existing).
Over to you!
This has been just a short introduction to the capabilities of VEP - have a look through
some more of the options, see them all on the command line using --help, or try using the
shortcut --everything which switches on almost all available output fields! Try out the
different options in the filtering tool, and if you're feeling adventurous why not use some of
your own data to annotate your variants or have a go with a plugin or two.
!
Variant Effect Predictor Download and
install
Download
Download ensembl-vep package (see below the different ways to download it) and then follow
the installation instructions.
Using Git
Clone the Git repository
Use git to download the ensembl-vep package:
git clone https://github.com/Ensembl/ensembl-vep.git
cd ensembl-vep
Update to a newer version
To update from a previous version:
cd ensembl-vep
git pull
git checkout release/112
perl INSTALL.pl
Use an older version
To use an older version (this example shows how to set up release 87):
cd ensembl-vep
git checkout release/87
perl INSTALL.pl
Download the Zipped package file
Users without the git utility installed may download a zip file from GitHub, though we would
always recommend using git if possible.
curl -L -O https://github.com/Ensembl/ensembl-
vep/archive/release/112.zip
unzip 112.zip
cd ensembl-vep-release-112/
Previous versions (ensembl-tools)
Previously VEP was available as part of the ensembl-tools package (see the Ensembl archive
site for documentation). The following downloads are available for archival purposes.
Show versions
What's new?
New in version 112 (January 2024)
Enhanced Structural Variant Support:

Added support for CNV:TR
Enabled the use of chromosome synonyms in breakends
Report consequences for each breakend and enable the input of single breakends
New plugins (supported on CLI, Web and REST):
AlphaMissense - uses a standardized catalog of human Ribo-seq ORFs to re-
calculate consequences for variants located in these translated regions
New plugins (supported on CLI and Web):
RiboseqORFs - uses a standardized catalog of human Ribo-seq ORFs to re-
calculate consequences for variants located in these translated regions
New plugins (supported on CLI):
Paralogues - fetches variants overlapping the genomic coordinates of amino acids
aligned between paralogue proteins
AVADA - Automatic VAriant evidence DAtabase is a novel machine learning tool
that uses natural language processing to automatically identify pathogenic genetic
variant evidence in full-text primary literature about monogenic disease and convert it
to genomic coordinates.
Plugin support added to REST and Web for:
CADD_SV
CADD scores for Sus scrofa
Dosage Sensitivity
Enformer
Previous version history - from version 88: Show
Older versions (ensembl-tools) - until version 87: Show
Requirements
VEP requires:
gcc, g++ and make
Perl version 5.10 or above recommended (tested on 5.10, 5.14, 5.18, 5.22, 5.26)
Perl packages:
Archive::Zip
DBD::mysql (version <=4.050)
DBI
See this guide for more information on how to install perl modules.
Additional libraries can be installed for extra features and enhancements but they are not
required to run VEP in most of the use cases.
VEP's INSTALL.pl script will install required components of Ensembl API for you, but VEP may
also be used with any pre-existing API installations you have, provided their versions match
the version of VEP you are using.
VEP has been developed for UNIX-like environments and works well on Linux (e.g. Ubuntu,
Debian, Mint) and Mac OSX.
It can also be used on Windows systems with a more involved installation process.
Installation

VEP's INSTALL.pl makes it easy to set up your environment for using the VEP. It will download
and configure a minimal set of the Ensembl API for use by the VEP, and can also download
cache files, FASTA files and plugins.
Run the following, and follow any prompts as they appear:
perl INSTALL.pl
Additional non-essential components and enhancements must be installed manually.
Software components installed
BioPerl
ensembl
ensembl-io
ensembl-variation
ensembl-funcgen
Bio::DB::HTS
If you already have the latest version of the API installed you do not need to run the installer,
although it can be used to simply update your API version (with post-release patches applied),
and retrieve cache and FASTA files. The installer downloads the API within the VEP directory and
will not affect any other Ensembl API installations.
The script will also attempt to install a Perl::XS module, Bio::DB::HTS , for rapid access to
bgzipped FASTA files. If this fails, you may add the --NO_HTSLIB flag when running the installer;
VEP will fall back to using Bio::DB::Fasta for this functionality (more details).
Running the installer
The installer is run on the command line as follows:
perl INSTALL.pl [options]
Follow on-screen prompts and note warnings of any files which will be deleted/overwritten
You should not need to add any options, but configuration of the installer is possible with the flags
below. Options can also be set by exporting environment variables prefixed with VEP_ before
running the installer (for instance, export VEP_NO_HTSLIB=1 and export
VEP_DIR_PLUGINS="/plugins").
Flag Alternate Description
--
ASSEM
BLY
-y
Assembly version to use when using --AUTO. Most species have only one
assembly available on each software release; currently this is only required
for human on release 76 onwards.
--
AUTO
-a
Run installer without prompts. Use the following options to specify parts to
install:
a (API + Bio::DB::HTS/htslib)
l (Bio::DB::HTS/htslib only)
c (cache)
f (FASTA)
p (plugins) — Require the use of the --PLUGINS flag to list the
plugin(s) to install.
e.g. for API and cache:
perl INSTALL.pl --AUTO ac

--
CACHE
_VERS
ION
[vers
ion]
! By default the installer will download the latest version of VEP caches and
FASTA files (currently 112). You can force the script to install a different
version, but there is no guarantee that a version of the API will be
compatible with a different version of the cache.
--
CACHE
DIR
[dir]
-c
By default the script will install the cache files in the ".vep" subdirectory in
your home area. This option configures where cache files are installed.
The --dir_cache flag must be passed when running the VEP if a non-default
cache directory is given:
./vep --dir_cache [dir]
--
DESTD
IR
[dir]
-d
By default the script will install the API modules in a subdirectory of the
current directory named "Bio". Using this option you can configure where
the Bio directory is created. If something other than the default is used, this
directory must either be added to your PERL5LIB environment variable
when running the VEP, or included using perl's -I flag:
perl -I [dir] vep
--
NO_HT
SLIB
-l
Don't attempt to install Bio::DB::HTS/htslib
--
NO_TE
ST
! Don't run API tests - useful if you know a harmless failure will prevent
continuation of the installer
--
NO_UP
DATE
-n
By default the script will check for new versions or updates of the VEP.
Using this option will skip this check.
--
PLUGI
NS
-g
Comma-separated list of plugins to install when using --AUTO. To install all
available plugins, use --PLUGINS all.
# List the available plugins:
perl INSTALL.pl -a p --PLUGINS list
# Download/install all the available plugins:
perl INSTALL.pl -a p --PLUGINS all
# Download/install a defined list of plugins, e.g.:
perl INSTALL.pl -a p --PLUGINS dbNSFP,CADD,G2P
--
PLUGI
NSDIR
[dir]
-r
By default the script will install the plugins files in the "Plugins" subdirectory
of the --CACHEDIR directory. This option configures where the plugins files
are installed.
The --dir_plugins flag must be passed when running the VEP if a non-
default plugins directory is given:
./vep --dir_plugins [dir]
--
PREFE
R_BIN
-p
Use this if the installer fails with out of memory errors.

--
SPECI
ES
-s
Comma-separated list of species to install when using --AUTO. To install
the RefSeq cache, add "_refseq" to the species name, e.g.
"homo_sapiens_refseq", or "_merged" to install the merged
Ensembl/RefSeq cache. Remember to use --refseq or --merged when
running the VEP with the relevant cache!
Use all to install data for all available species.
--
QUIET
-q
Don't write any status output when using --AUTO.
Additional components
INSTALL.pl will set up the minimum requirements for VEP. Some features and enhancements,
however, require the installation of additional components. Most are perl modules that are easily
installed using cpanm; see this guide for more information on how to install perl modules.
Typically, you will use cpanm to install modules locally in your home directories; this shows how
to set up a path for perl modules and install one there:
mkdir -p $HOME/cpanm
export PERL5LIB=$PERL5LIB:$HOME/cpanm/lib/perl5
cpanm -l $HOME/cpanm Set::IntervalTree
To make the change to PERL5LIB permanent, it is recommended to add the export line to your
$HOME/.bashrc or $HOME/.profile.
Additional features
JSON - required to produce JSON format output
Set::IntervalTree - used to find overlaps between entities in coordinate space.
Required to use --nearest
Bio::DB::BigFile - required to use bigWig format custom annotation files. See
Bio::DB::BigFile instructions.
Speed enhancements - these modules can improve VEP runtime
PerlIO::gzip - marginal gains in compressed file parsing as used by VEP cache
ensembl-xs - provides pre-compiled replacements for frequently used routines in VEP.
Requires manual installation, see README for details
Bio::DB::BigFile
In order for VEP to be able to access bigWig format custom annotation files, the Bio::DB::BigFile
perl module is required. Installation involves downloading and compiling the kent source tree .
The current version of the kent source tree does not work correctly with Bio::DB::BigFile, so it is
necessary to install an archive version known to work (v335).
1.
Download and unpack the kent source tree
wget
https://github.com/ucscGenomeBrowser/kent/archive/v335_base.tar.gz
tar xzf v335_base.tar.gz
2.
Set up some environment variables; these are required only temporarily for this installation
process
export KENT_SRC=$PWD/kent-335_base/src
export MACHTYPE=$(uname -m)
export CFLAGS="-fPIC"
export MYSQLINC=`mysql_config --include | sed -e 's/^-I//g'`
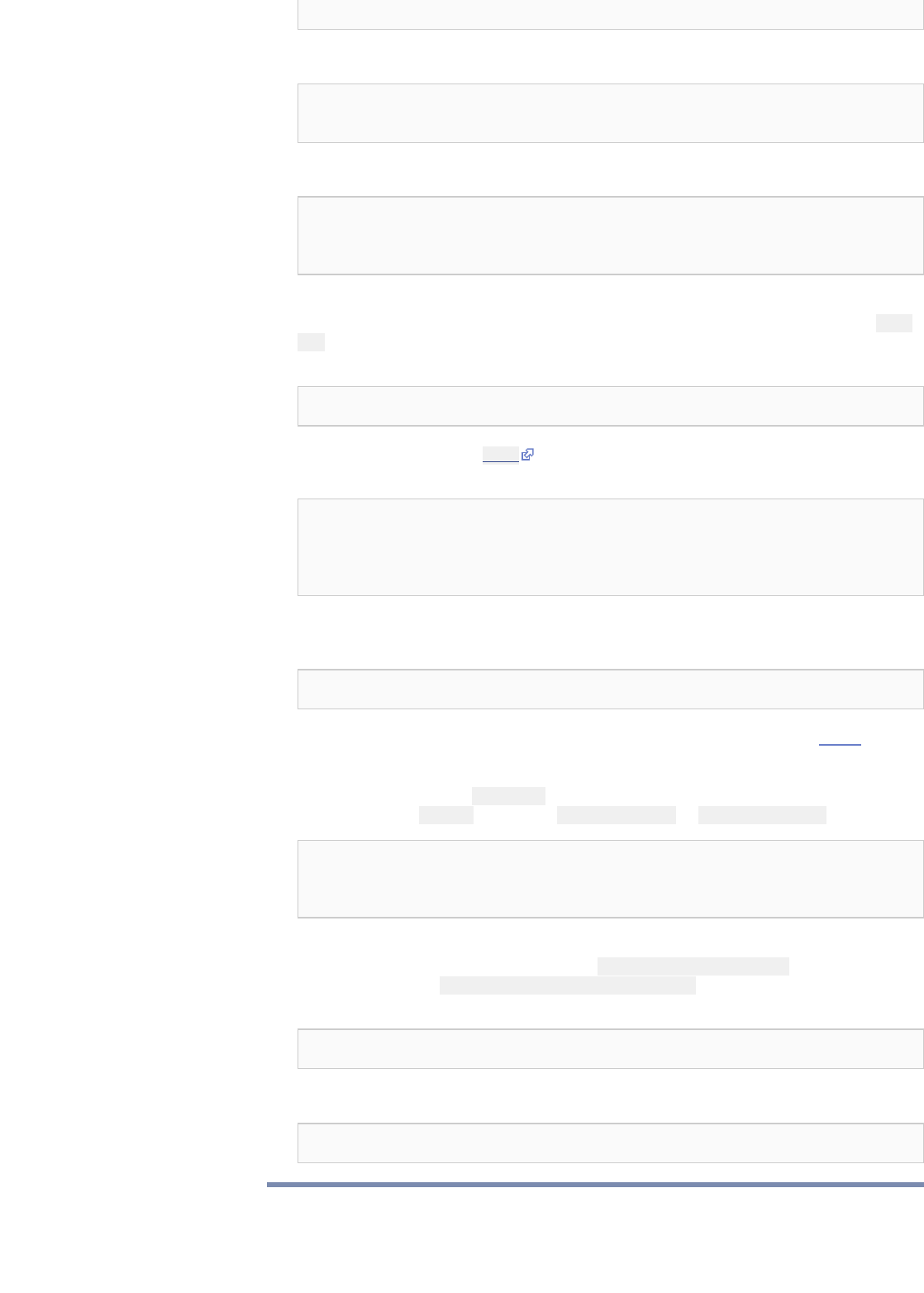
export MYSQLLIBS=`mysql_config --libs`
3.
Modify kent build parameters
cd $KENT_SRC/lib
echo 'CFLAGS="-fPIC"' > ../inc/localEnvironment.mk
4.
Build kent source
make clean && make
cd ../jkOwnLib
make clean && make
If either of these steps fail, you may have some missing dependencies. Known common
missing dependencies are libpng and libssl; these may be installed, for example, with apt-
get on Ubuntu. If you do not have sudo access you may have to ask your sysadmin to
install any missing dependencies.
sudo apt-get install libpng-dev libssl-dev
On Mac OSX you may use brew ; the openssl libraries also need to be symbolically linked
to a different path:
brew install libpng openssl
cd /usr/local/include
ln -s ../opt/openssl/include/openssl .
cd -
5.
On some systems (e.g. Mac OSX), a compiled file is placed in a path that Bio::DB::BigFile
cannot find. You can correct this with:
ln -s $KENT_SRC/lib/x86_64/* $KENT_SRC/lib/
6.
We'll now use cpanm to install the perl module for Bio::DB::BigFile itself. See above for
guidance on this. In this example we're going to install the module to a path within your
home directory. In order to do this we must modify the paths that perl looks in to find
modules by adding to the PERL5LIB environment module. To make this change permanent
you must add the export line to your $HOME/.bashrc or $HOME/.profile.
mkdir -p $HOME/cpanm
export PERL5LIB=$PERL5LIB:$HOME/cpanm/lib/perl5
cpanm -l $HOME/cpanm Bio::DB::BigFile
If you are prompted for the path to the kent source tree, that means something didn't go right
in the compilation above. Double check that $KENT_SRC/lib/jkweb.a exists and is not
found instead at e.g. $KENT_SRC/lib/x86_64/jkweb.a. You may copy or link the file
(and the other files in that directory) to the former path.
ln -s $KENT_SRC/lib/x86_64/* $KENT_SRC/lib/
7.
You should now be able to successfully run the appropriate test in the VEP package:
perl -Imodules t/AnnotationSource_File_BigWig.t
Using VEP in Mac OS
Installing VEP on Mac OS is slightly trickier than other Linux-based systems, and will require
additional dependancies.

These instructions will guide you through the setup of Perlbrew, Homebrew, MySQL and other
dependancies that will allow for a clean installation of VEP on your Mac OS system.
These instructions have been tested on macOS High Sierra (10.13) and macOS Sierra (10.12).
Older versions may require additional tweaks, however we shall endeavor to keep these
instructions up to date for future versions of MacOS.
Prerequisite Setup
List of prerequisites: XCode, GCC, Perlbrew, Cpanm, Homebrew, mysql, DBI, DBD::mysql
(version <=4.050)
XCode and GCC
VEP requires XCode and GCC for installation purposes. Fortunately, recent versions of macOS
will look for (and attempt to install if required) both of these when you run the following command:
gcc -v
Perlbrew
We recommend using Perlbrew to install a new version of Perl on your mac, to prevent messing
with the vendor perl too much. This can be done with the following command:
curl -L http://install.perlbrew.pl | bash
echo 'source $HOME/perl5/perlbrew/etc/bashrc' >> ~/.bash_profile
At this point, PLEASE RESTART YOUR TERMINAL WINDOW to allow for the perlbrew changes
to take effect.
We recommend installing Perl version 5.26.2 to run VEP, and installing cpanm to handle the
installation of perl modules.
These steps can be completed with the commands:
perlbrew install -j 5 --as 5.26.2 --thread --64all -Duseshrplib
perl-5.26.2 --notest
perlbrew switch 5.26.2
perlbrew install-cpanm
Homebrew
This package management system for Mac OS would make the installation of the next
prerequisite (i.e. xs) easier.
/usr/bin/ruby -e "$(curl -fsSL
https://raw.githubusercontent.com/Homebrew/install/master/install)"
xz
VEP requires the installation of xz, a data-compression utility. The easiest way to install the xz
package is through homebrew:
brew install xz
MySQL
In order to connect to the Ensembl databases, a collection of MySQL related dependancies are
required. Fortunately, these can be installed neatly with Homebrew and Cpanm:
brew install mysql
cpanm DBI
cpanm DBD::mysql@4.050

Installing BioPerl
On some versions of macOS, the VEP installer fails to cleanly install BioPerl, so a manual install
will prevent issues:
curl -O
https://cpan.metacpan.org/authors/id/C/CJ/CJFIELDS/BioPerl-1.6.924.ta
r.gz
tar zxvf BioPerl-1.6.924.tar.gz
echo 'export PERL5LIB=${PERL5LIB}:##PATH_TO##/bioperl-1.6.924' >>
~/.bash_profile
where ##PATH_TO##/bioperl-1.6.924 refers to the location of the newly unzipped BioPerl
directory.
Final Dependancies
Installing the following Perl modules with cpanm will allow for full VEP functionality:
cpanm Test::Differences Test::Exception Test::Perl::Critic
Archive::Zip PadWalker Error Devel::Cycle Role::Tiny::With
Module::Build
export DYLD_LIBRARY_PATH=/usr/local/mysql/lib/:$DYLD_LIBRARY_PATH
Installing VEP
And that should be that! You should now be able to install VEP using the installer:
git clone https://github.com/ensembl/ensembl-vep
cd ensembl-vep
perl INSTALL.pl --NO_TEST
Using VEP in Windows
VEP was developed as a command-line tool, and as a Perl script its natural environment is a
Linux system. However, there are several ways you can use VEP on a Windows machine.
You may also consider using VEP's web or REST interfaces.
Virtual machines
Using a virtual machine you can run a virtual Linux system in a window on your machine. There
are two ways to do this:
1.
Use the Ensembl virtual machine image
2.
Use Docker
Perl
If Perl is installed on Windows, VEP can be setup. However this may require installation of
dependent modules. We recommend using Docker to run VEP on Windows.
1.
Check Perl is installed
2.
Download and unpack the zip of the ensembl-vep package
3.
Open a Command Prompt (search for Command Prompt in the Start Menu)
4.
Navigate to the directory where you unpacked the VEP package, e.g.

cd Downloads/ensembl-vep-release-112
5.
Run INSTALL.pl with --NO_HTSLIB and --NO_TEST; you will see some warnings about the
"which" command not being available (these will also appear when running VEP and can be
ignored).
perl INSTALL.pl --NO_HTSLIB --NO_TEST
Docker
Docker allows running applications in virtualised containers. The VEP Docker image is
available from DockerHub: VEP in DockerHub
After installing Docker , download the VEP Docker image:
docker pull ensemblorg/ensembl-vep
To download cache files and other data with VEP Docker, we recommend mounting a directory
from your local (host) machine to folder /data from the Docker image. For instance:
mkdir $HOME/vep_data
docker run -t -i -v $HOME/vep_data:/data ensemblorg/ensembl-vep
In the example above, data in $HOME/vep_data will be accessible by both the local machine
and VEP Docker. The Ensembl VEP API, plugins and their dependencies (e.g. Perl APIs,
Bio::DB::HTS, htslib, ...) are already installed in the image.
Cache and FASTA files installation
You can run the INSTALL.pl script to install the cache and FASTA files:
docker run -t -i -v $HOME/vep_data:/data ensemblorg/ensembl-vep
INSTALL.pl
You will be asked to install cache data. Type the comma-separated numbers for the
species/assembly of interest and press enter. Your data will download and unpack; this
may take a while.
If you wish to retrieve HGVS annotations, please download the FASTA files for your species.
To do this, at the next prompt type 0 and press enter.
The above process may also be performed in one command; for example, to set up the cache
and corresponding FASTA for human GRCh38:
docker run -t -i -v $HOME/vep_data:/data ensemblorg/ensembl-vep
INSTALL.pl -a cf -s homo_sapiens -y GRCh38
The installer downloads VEP data to the mounted directory (e.g., $HOME/vep_data). The
downloaded data will be automatically detected as long as its folder is mounted when running
VEP:
docker run -v $HOME/vep_data:/data ensemblorg/ensembl-vep vep -i
examples/homo_sapiens_GRCh38.vcf --cache
Running VEP with data from local folder
Here is an example on running VEP with data from folder $HOME/vep_data in the local machine

(provided that the cache has been downloaded to that folder):
docker run -v $HOME/vep_data:/data ensemblorg/ensembl-vep \
vep --cache --offline --format vcf --vcf --force_overwrite \
--input_file input/my_input.vcf \
--output_file output/my_output.vcf \
--custom
file=custom/my_extra_data.bed,short_name=BED_DATA,format=bed,type=exa
ct,coords=1 \
--plugin NMD
Please avoid using absolute paths to data as the paths inside the container differ from your local
machine.
Update from a previous version
1.
Update your Docker container
docker pull ensemblorg/ensembl-vep
2.
Update your cache
# Install the new cache through the VEP INSTALL.pl script (see
"Cache installation" section above)
docker run -t -i -v $HOME/vep_data:/data ensemblorg/ensembl-vep
INSTALL.pl -a c
# Or install the cache manually
cd $HOME/vep_data
curl -O
https://ftp.ensembl.org/pub/release-112/variation/vep/homo_sapiens
_vep_112_GRCh38.tar.gz
tar xzf homo_sapiens_vep_112_GRCh38.tar.gz
Singularity
Due to root requirements for the Docker daemon, using the Docker container for VEP is not
always possible to HPC users. Singularity, an alternative containerisation tool, does not assume
that you have a system where you are the root user. This has led to increased popularity in HPC
contexts due to increased access rights flexibility.
After installing Singularity , VEP may be used with Singularity based on the VEP Docker image
from DockerHub:
singularity pull --name vep.sif docker://ensemblorg/ensembl-vep
The following is a brief example showing how to use a directory on your local (host) machine to
store cache data for VEP.
mkdir $HOME/vep_data
singularity exec vep.sif vep --dir $HOME/vep_data --help
The Ensembl VEP API, plugins and their dependencies (e.g. Perl APIs, Bio::DB::HTS, htslib, ...)
are already installed in the image.
Cache and FASTA files installation
You can run the INSTALL.pl script to install the Cache data and FASTA files. For example, to set
up the cache and corresponding FASTA for human GRCh38 in your local folder
$HOME/vep_data:

!
singularity exec vep.sif INSTALL.pl -c $HOME/vep_data -a cf -s
homo_sapiens -y GRCh38
The installer downloads data to the specified directory (e.g., $HOME/vep_data). When running
VEP via Singularity, point to this directory using --dir:
singularity exec vep.sif vep --dir $HOME/vep_data -i
examples/homo_sapiens_GRCh38.vcf --cache
Running VEP with data from local folder
Here is an example on running VEP with data from folder $HOME/vep_data in the local machine
(provided that the cache has been downloaded to that folder):
singularity exec vep.sif \
vep --dir $HOME/vep_data \
--cache --offline --format vcf --vcf --force_overwrite \
--input_file input/my_input.vcf \
--output_file output/my_output.vcf \
--custom
file=custom/my_extra_data.bed,short_name=BED_DATA,format=bed,type=exa
ct,coords=1 \
--plugin NMD
Update from a previous version
1.
Update your docker container
singularity pull --name vep.sif docker://ensemblorg/ensembl-vep
2.
Update your cache
# Install the new cache through the VEP INSTALL.pl script (see
"Cache installation" section above)
singularity exec vep.sif INSTALL.pl -c $HOME/vep_data -a c
# Or install the cache manually
cd $HOME/vep_data
curl -O
https://ftp.ensembl.org/pub/release-112/variation/vep/homo_sapiens
_vep_112_GRCh38.tar.gz
tar xzf homo_sapiens_vep_112_GRCh38.tar.gz

Variant Effect Predictor Data formats
Input
Both the web and script version of VEP can use the same input formats. Formats can be auto-detected by the VEP script, but must be
manually selected when using the web interface.
VEP can use different input formats:
Format Variant example Structural variant example
Default VEP input 1 881907 881906 -/C + 1 160283 471362 DUP +
VCF 1 65568 . A C . . . 1 7936271 . N N[12:58877476[ . . SVTYPE=BND
HGVS identifiers ENST00000618231.3:c.9G>C ✗ Not supported
Variant identifiers rs699 nsv1000164
Genomic SPDI notation NC_000016.10:68684738:G:A ✗ Not supported
REST-style regions 14:19584687-19584687:-1/T 21:25587759-25587769/DEL
Default VEP input
The default format is a simple whitespace-separated format (columns may be separated by space or tab characters), containing five
required columns plus an optional identifier column:
1.
chromosome - just the name or number, with no 'chr' prefix
2.
start
3.
end
4.
allele - pair of alleles separated by a '/', with the reference allele first (or structural variant type)
5.
strand - defined as + (forward) or - (reverse). The strand will only be used for VEP to know which alleles to use.
6.
identifier - this identifier will be used in VEP's output. If not provided, VEP will construct an identifier from the given coordinates and
alleles.
1 881907 881906 -/C +
2 946507 946507 G/C +
5 140532 140532 T/C +
8 150029 150029 A/T + var2
12 1017956 1017956 T/A +
14 19584687 19584687 C/T -
19 66520 66520 G/A + var1
An insertion (of any size) is indicated by start coordinate = end coordinate + 1. For example, an insertion of 'C' between nucleotides
12600 and 12601 on the forward strand of chromosome 8 is indicated as follows:
8 12601 12600 -/C +
A deletion is indicated by the exact nucleotide coordinates. For example, a three base pair deletion of nucleotides 12600, 12601, and
12602 of the reverse strand of chromosome 8 will be:
8 12600 12602 CGT/- -
Structural variants are also supported by indicating a structural variant type instead of the allele:
1 20000 30000 CN4 + cnv4
1 160283 471362 DUP + dup

1 1385015 1387562 DEL + del1
12 1017956 1017956 INV + inv
21 25587759 25587769 CN0 + del2
VCF
VEP also supports using VCF (Variant Call Format) version 4.0 . This is a common format used by the 1000 genomes project, and can
be produced as an output format by many variant calling tools:
#CHROM POS ID REF ALT QUAL FILTER INFO FORMAT
1 65568 . A C . . . .
1 230710048 rs699 A G . . . .
2 265023 . C T . . . .
3 319780 . GA G . . . .
20 3 . C CAAG,CAAGAAG . PASS . .
21 43762120 rs1300 T A,C,G . . . .
Structural variants are also supported depending on structural variant type.
Users using VCF should note a peculiarity in the difference between how Ensembl and VCF describe unbalanced variants. For any
unbalanced variant (i.e. insertion, deletion or unbalanced substitution), the VCF specification requires that the base immediately before
the variant should be included in both the reference and variant alleles. This also affects the reported position i.e. the reported position
will be one base before the actual site of the variant.
In order to parse this correctly, VEP needs to convert such variants into Ensembl-type coordinates, and it does this by removing the
additional base and adjusting the coordinates accordingly. This means that if an identifier is not supplied for a variant (in the 3rd column
of the VCF), then the identifier constructed and the position reported in VEP's output file will differ from the input.
This problem can be overcome with the following:
1.
ensuring each variant has a unique identifier specified in the 3rd column of the VCF
2.
using VCF format as output (--vcf) - this preserves the formatting of your input coordinates and alleles
3.
using --minimal and --allele_number (see Complex VCF entries).
The following examples illustrate how VCF describes a variant and how it is handled internally by VEP. Consider the following aligned
sequences (for the purposes of discussion on chromosome 20):
Ref: a t C g a // C is the reference base
1 : a t G g a // C base is a G in individual 1
2 : a t - g a // C base is deleted w.r.t. the reference in individual 2
3 : a t CAg a // A base is inserted w.r.t. the reference sequence in individual 3
Individual 1
The first individual shows a simple balanced substitution of G for C at base 3. This is described in a compatible manner in VCF and
Ensembl styles. Firstly, in VCF:
20 3 . C G . PASS .
And in Ensembl format:
20 3 3 C/G +
Individual 2
The second individual has the 3rd base deleted relative to the reference. In VCF, both the reference and variant allele columns must
include the preceding base (T) and the reported position is that of the preceding base:
20 2 . TC T . PASS .

In Ensembl format, the preceding base is not included, and the start/end coordinates represent the region of the sequence deleted. A "-"
character is used to indicate that the base is deleted in the variant sequence:
20 3 3 C/- +
The upshot of this is that while in the VCF input file the position of the variant is reported as 2, in the output file from VEP the position will
be reported as 3. If no identifier is provided in the third column of the VCF, then the constructed identifier will be:
20_3_C/-
Individual 3
The third individual has an "A" inserted between the 3rd and 4th bases of the sequence relative to the reference. In VCF, as for the
deletion, the base before the insertion is included in both the reference and variant allele columns, and the reported position is that of the
preceding base:
20 3 . C CA . PASS .
In Ensembl format, again the preceding base is not included, and the start/end positions are "swapped" to indicate that this is an
insertion. Similarly to a deletion, a "-" is used to indicate no sequence in the reference:
20 4 3 -/A +
Again, the output will appear different, and the constructed identifier may not be what is expected:
20_3_-/A
Using VCF format output, or adding unique identifiers to the input (in the third VCF column), can mitigate this issue.
Complex VCF entries
For VCF entries with multiple alternate alleles, VEP will only trim the leading base from alleles if all REF and ALT alleles start with the
same base:
20 3 . C CAAG,CAAGAAG . PASS .
This will be considered internally by VEP as equivalent to:
20 4 3 -/AAG/AAGAAG +
Now consider the case where a single VCF line contains a representation of both a SNV and an insertion:
20 3 . C CAAAG,G . PASS .
Here the input alleles will remain unchanged, and VEP will consider the first REF/ALT pair as a substitution of C for CAAG, and the
second as a C/G SNV:
20 3 3 C/CAAG/G +
To modify this behaviour, VEP script users may use --minimal. This flag forces VEP to consider each REF/ALT pair independently,
trimming identical leading and trailing bases from each as appropriate. Since this can lead to confusing output regarding coordinates etc,
it is not the default behaviour. It is recommended to use the --allele_number flag to track the correspondence between alleles as input
and how they appear in the output.
Structural variant types

VEP can also call consequences on structural variants using the following input formats:
Default VEP input
REST-style regions
Variant identifiers
VCF
To recognise a variant as a structural variant, the allele string (or SVTYPE in the INFO column of the VCF format) must be set to one of
the currently supported values:
INS - insertion
DEL - deletion
DUP - duplication
TDUP - tandem duplication
INV - inversion
CNV - copy number variation
The copy number value can be specified, such as <CN0> or <CN=4>
BND - breakend
In VCF, breakend replacements are inserted into the ALT column and need to meet the HTS specifications , such as
A[22:22893780[,A[X:10932343[.
Examples of structural variants encoded in VCF format:
#CHROM POS ID REF ALT QUAL FILTER INFO
1 160283 dup . <DUP> . . SVTYPE=DUP;END=471362
1 1385015 del . <DEL> . . SVTYPE=DEL;END=1387562
1 7936271 bnd N N[12:58877476[ . . SVTYPE=BND
See the VCF definition document for more detail on how to describe structural variants in VCF format.
HGVS identifiers
See https://varnomen.hgvs.org for details. These must be relative to genomic or Ensembl transcript coordinates.
It also is possible to use RefSeq transcripts in both the web interface and the VEP script (see script documentation): this works for
RefSeq transcripts that align to the genome correctly.
Examples:
ENST00000618231.3:c.9G>C
ENST00000471631.1:c.28_33delTCGCGG
ENST00000285667.3:c.1047_1048insC
5:g.140532G>C
Examples using RefSeq identifiers (using --refseq in the VEP script, or select the otherfeatures transcript database on the web interface
and input type of HGVS):
NM_153681.2:c.7C>T
NM_005239.6:c.190G>A
NM_001025204.2:c.336G>A
HGVS protein notations may also be used, provided that they unambiguously map to a single genomic change. Due to redundancy in the
amino acid code, it is not always possible to work out the corresponding genomic sequence change for a given protein sequence
change. The following example is for a permissable protein notation in dog (Canis familiaris):
ENSCAFP00000040171.1:p.Thr92Asn

Ambiguous gene-based descriptions
It is possible to use ambiguous descriptions listing only gene symbol or UniProt accession and protein change (e.g.
PHF21B:p.Tyr124Cys, P01019:p.Ala268Val), as seen in the literature, though this is not recommended as it can produce multiple
different variants at genomic level. The transcripts for a gene are considered in the following order:
1.
MANE Select transcript status
2.
MANE Plus Clinical transcript status
3.
canonical status of transcript
4.
APPRIS isoform annotation
5.
transcript support level
6.
biotype of transcript ("protein_coding" preferred)
7.
CCDS status of transcript
8.
consequence rank according to this table
9.
translated, transcript or feature length (longer preferred)
and the first compatible transcript is used to map variants to the genome for annotation.
Variant identifiers
These should be dbSNP rsIDs (such as rs699), or any synonym for a variant present in the Ensembl Variation database. Structural
variant identifiers (like nsv1000164 and esv1850194) are also supported.
See here for a list of identifier sources in Ensembl.
Examples:
rs1156485833
rs1258750482
rs867704559
esv1815690
nsv1000164
Genomic SPDI notation
VEP can also support genomic SPDI notation which uses four fields delimited by colons S:P:D:I (Sequence:Position:Deletion:Insertion).
In SPDI notation, the position refers to the base before the variant, not the base of the variant itsef.
See here for details.
Examples:
NC_000016.10:68684738:G:A
NC_000017.11:43092199:GCTTTT:
NC_000013.11:32315789::C
NC_000016.10:68644746:AA:GTA
16:68684738:2:AC
REST-style regions
VEP's region REST endoint requires variants are described as [chr]:[start]-[end]:[strand]/[allele].

This follows the same conventions as the default input format, with the key difference being that this format does not require the
reference (REF) allele to be included; VEP will look up the reference allele using either a provided FASTA file (preferred) or Ensembl
core database. Strand is optional and defaults to 1 (forward strand).
# SNP
5:140532-140532:1/C
# SNP (reverse strand)
14:19584687-19584687:-1/T
# insertion
1:881907-881906:1/C
# 5bp deletion
2:946507-946511:1/-
Structural variants are also supported by indicating a structural variant type in the place of the [allele]:
# structural variant: deletion
21:25587759-25587769/DEL
# structural variant: inversion
21:25587759-25587769/INV
Output
VEP can return the results in different formats:
Default VEP output
Tab-delimited output
VCF
JSON output
Along with the results VEP computes and returns some statistics.
Default VEP output
The default output format ("VEP" format when downloading from the web interface) is a 14 column tab-delimited file. Empty values are
denoted by '-'. The output columns are:
1.
Uploaded variation - as chromosome_start_alleles
2.
Location - in standard coordinate format (chr:start or chr:start-end)
3.
Allele - the variant allele used to calculate the consequence
4.
Gene - Ensembl stable ID of affected gene
5.
Feature - Ensembl stable ID of feature
6.
Feature type - type of feature. Currently one of Transcript, RegulatoryFeature, MotifFeature.
7.
Consequence - consequence type of this variant
8.
Position in cDNA - relative position of base pair in cDNA sequence
9.
Position in CDS - relative position of base pair in coding sequence
10.
Position in protein - relative position of amino acid in protein
11.
Amino acid change - only given if the variant affects the protein-coding sequence
12.
Codon change - the alternative codons with the variant base in upper case

13.
Co-located variation - identifier of any existing variants. Switch on with --check_existing
14.
Extra - this column contains extra information as key=value pairs separated by ";", see below.
Other output fields:
REF_ALLELE - the reference allele (after minimisation)
UPLOADED_ALLELE - the uploaded allele string (before minimisation)
IMPACT - the impact modifier for the consequence type
VARIANT_CLASS - Sequence Ontology variant class
SYMBOL - the gene symbol
SYMBOL_SOURCE - the source of the gene symbol
STRAND - the DNA strand (1 or -1) on which the transcript/feature lies
ENSP - the Ensembl protein identifier of the affected transcript
FLAGS - transcript quality flags:
cds_start_NF: CDS 5' incomplete
cds_end_NF: CDS 3' incomplete
SWISSPROT - Best match UniProtKB/Swiss-Prot accession of protein product
TREMBL - Best match UniProtKB/TrEMBL accession of protein product
UNIPARC - Best match UniParc accession of protein product
HGVSc - the HGVS coding sequence name
HGVSp - the HGVS protein sequence name
HGVSg - the HGVS genomic sequence name
HGVS_OFFSET - Indicates by how many bases the HGVS notations for this variant have been shifted. Value must be greater than
0.
NEAREST - Identifier(s) of nearest transcription start site
SIFT - the SIFT prediction and/or score, with both given as prediction(score)
PolyPhen - the PolyPhen prediction and/or score
MOTIF_NAME - the source and identifier of a transcription factor binding profile aligned at this position
MOTIF_POS - The relative position of the variation in the aligned TFBP
HIGH_INF_POS - a flag indicating if the variant falls in a high information position of a transcription factor binding profile (TFBP)
MOTIF_SCORE_CHANGE - The difference in motif score of the reference and variant sequences for the TFBP
CELL_TYPE - List of cell types and classifications for regulatory feature
CANONICAL - a flag indicating if the transcript is denoted as the canonical transcript for this gene
CCDS - the CCDS identifer for this transcript, where applicable
INTRON - the intron number (out of total number)
EXON - the exon number (out of total number)
DOMAINS - the source and identifer of any overlapping protein domains
DISTANCE - Shortest distance from variant to transcript
IND - individual name
ZYG - zygosity of individual genotype at this locus
SV - IDs of overlapping structural variants
FREQS - Frequencies of overlapping variants used in filtering
AF - Frequency of existing variant in 1000 Genomes
AFR_AF - Frequency of existing variant in 1000 Genomes combined African population
AMR_AF - Frequency of existing variant in 1000 Genomes combined American population
ASN_AF - Frequency of existing variant in 1000 Genomes combined Asian population

EUR_AF - Frequency of existing variant in 1000 Genomes combined European population
EAS_AF - Frequency of existing variant in 1000 Genomes combined East Asian population
SAS_AF - Frequency of existing variant in 1000 Genomes combined South Asian population
gnomADe_AF - Frequency of existing variant in gnomAD exomes combined population
gnomADe_AFR_AF - Frequency of existing variant in gnomAD exomes African/American population
gnomADe_AMR_AF - Frequency of existing variant in gnomAD exomes American population
gnomADe_ASJ_AF - Frequency of existing variant in gnomAD exomes Ashkenazi Jewish population
gnomADe_EAS_AF - Frequency of existing variant in gnomAD exomes East Asian population
gnomADe_FIN_AF - Frequency of existing variant in gnomAD exomes Finnish population
gnomADe_NFE_AF - Frequency of existing variant in gnomAD exomes Non-Finnish European population
gnomADe_OTH_AF - Frequency of existing variant in gnomAD exomes combined other combined populations
gnomADe_SAS_AF - Frequency of existing variant in gnomAD exomes South Asian population
gnomADg_AF - Frequency of existing variant in gnomAD exomes combined population
gnomADg_AFR_AF - Frequency of existing variant in gnomAD genomes African/American population
gnomADg_AMI_AF - Frequency of existing variant in gnomAD genomes Amish population
gnomADg_AMR_AF - Frequency of existing variant in gnomAD genomes American population
gnomADg_ASJ_AF - Frequency of existing variant in gnomAD genomes Ashkenazi Jewish population
gnomADg_EAS_AF - Frequency of existing variant in gnomAD genomes East Asian population
gnomADg_FIN_AF - Frequency of existing variant in gnomAD genomes Finnish population
gnomADg_MID_AF - Frequency of existing variant in gnomAD genomes Mid-eastern population
gnomADg_NFE_AF - Frequency of existing variant in gnomAD genomes Non-Finnish European population
gnomADg_OTH_AF - Frequency of existing variant in gnomAD genomes combined other combined populations
gnomADg_SAS_AF - Frequency of existing variant in gnomAD genomes South Asian population
MAX_AF - Maximum observed allele frequency in 1000 Genomes, ESP and gnomAD
MAX_AF_POPS - Populations in which maximum allele frequency was observed
CLIN_SIG - ClinVar clinical significance of the dbSNP variant
BIOTYPE - Biotype of transcript or regulatory feature
APPRIS - Annotates alternatively spliced transcripts as primary or alternate based on a range of computational methods. NB: not
available for GRCh37
TSL - Transcript support level. NB: not available for GRCh37
PUBMED - Pubmed ID(s) of publications that cite existing variant
SOMATIC - Somatic status of existing variant(s); multiple values correspond to multiple values in the Existing_variation field
PHENO - Indicates if existing variant is associated with a phenotype, disease or trait; multiple values correspond to multiple values
in the Existing_variation field
GENE_PHENO - Indicates if overlapped gene is associated with a phenotype, disease or trait
ALLELE_NUM - Allele number from input; 0 is reference, 1 is first alternate etc
MINIMISED - Alleles in this variant have been converted to minimal representation before consequence calculation
PICK - indicates if this block of consequence data was picked by --flag_pick or --flag_pick_allele
BAM_EDIT - Indicates success or failure of edit using BAM file
GIVEN_REF - Reference allele from input
USED_REF - Reference allele as used to get consequences
REFSEQ_MATCH - the RefSeq transcript match status; contains a number of flags indicating whether this RefSeq transcript
matches the underlying reference sequence and/or an Ensembl transcript (more information).
rseq_3p_mismatch: signifies a mismatch between the RefSeq transcript and the underlying primary genome assembly
sequence. Specifically, there is a mismatch in the 3' UTR of the RefSeq model with respect to the primary genome assembly
(e.g. GRCh37/GRCh38).
rseq_5p_mismatch: signifies a mismatch between the RefSeq transcript and the underlying primary genome assembly

sequence. Specifically, there is a mismatch in the 5' UTR of the RefSeq model with respect to the primary genome assembly.
rseq_cds_mismatch: signifies a mismatch between the RefSeq transcript and the underlying primary genome assembly
sequence. Specifically, there is a mismatch in the CDS of the RefSeq model with respect to the primary genome assembly.
rseq_ens_match_cds: signifies that for the RefSeq transcript there is an overlapping Ensembl model that is identical across the
CDS region only. A CDS match is defined as follows: the CDS and peptide sequences are identical and the genomic
coordinates of every translatable exon match. Useful related attributes are: rseq_ens_match_wt and rseq_ens_no_match.
rseq_ens_match_wt: signifies that for the RefSeq transcript there is an overlapping Ensembl model that is identical across the
whole transcript. A whole transcript match is defined as follows: 1) In the case that both models are coding, the transcript, CDS
and peptide sequences are all identical and the genomic coordinates of every exon match. 2) In the case that both transcripts
are non-coding the transcript sequences and the genomic coordinates of every exon are identical. No comparison is made
between a coding and a non-coding transcript. Useful related attributes are: rseq_ens_match_cds and rseq_ens_no_match.
rseq_ens_no_match: signifies that for the RefSeq transcript there is no overlapping Ensembl model that is identical across
either the whole transcript or the CDS. This is caused by differences between the transcript, CDS or peptide sequences or
between the exon genomic coordinates. Useful related attributes are: rseq_ens_match_wt and rseq_ens_match_cds.
rseq_mrna_match: signifies an exact match between the RefSeq transcript and the underlying primary genome assembly
sequence (based on a match between the transcript stable id and an accession in the RefSeq mRNA file). An exact match
occurs when the underlying genomic sequence of the model can be perfectly aligned to the mRNA sequence post polyA
clipping.
rseq_mrna_nonmatch: signifies a non-match between the RefSeq transcript and the underlying primary genome assembly
sequence. A non-match is deemed to have occurred if the underlying genomic sequence does not have a perfect alignment to
the mRNA sequence post polyA clipping. It can also signify that no comparison was possible as the model stable id may not
have had a corresponding entry in the RefSeq mRNA file (sometimes happens when accessions are retired or changed). When
a non-match occurs one or several of the following transcript attributes will also be present to provide more detail on the nature
of the non-match: rseq_5p_mismatch, rseq_cds_mismatch, rseq_3p_mismatch, rseq_nctran_mismatch, rseq_no_comparison
rseq_nctran_mismatch: signifies a mismatch between the RefSeq transcript and the underlying primary genome assembly
sequence. This is a comparison between the entire underlying genomic sequence of the RefSeq model to the mRNA in the
case of RefSeq models that are non-coding.
rseq_no_comparison: signifies that no alignment was carried out between the underlying primary genome assembly sequence
and a corresponding RefSeq mRNA. The reason for this is generally that no corresponding, unversioned accession was found
in the RefSeq mRNA file for the transcript stable id. This sometimes happens when accessions are retired or replaced. A
second possibility is that the sequences were too long and problematic to align (though this is rare).
OverlapBP - Number of base pairs overlapping with the corresponding structural variation feature
OverlapPC - Percentage of corresponding structural variation feature overlapped by the given input
CHECK_REF - Reports variants where the input reference does not match the expected reference
AMBIGUITY - IUPAC allele ambiguity code
Example of VEP default output format:
11_224088_C/A 11:224088 A ENSG00000142082 ENST00000525319 Transcript
missense_variant 742 716 239 T/N aCc/aAc - SIFT=deleterious(0);PolyPhen=unknown(0)
11_224088_C/A 11:224088 A ENSG00000142082 ENST00000534381 Transcript
5_prime_UTR_variant - - - - - - -
11_224088_C/A 11:224088 A ENSG00000142082 ENST00000529055 Transcript
downstream_variant - - - - - - -
11_224585_G/A 11:224585 A ENSG00000142082 ENST00000529937 Transcript
intron_variant - - - - - - HGVSc=ENST00000529937.1:c.136-346G>A
22_16084370_G/A 22:16084370 A - ENSR00000615113 RegulatoryFeature
regulatory_region_variant - - - - - - -
The VEP script will also add a header to the output file. This contains information about the databases connected to, and also a key
describing the key/value pairs used in the extra column.
## ENSEMBL VARIANT EFFECT PREDICTOR v112.0
## Output produced at 2017-03-21 14:51:27
## Connected to homo_sapiens_core_112_38 on ensembldb.ensembl.org
## Using cache in /homes/user/.vep/homo_sapiens/112_GRCh38
## Using API version 112, DB version 112
## polyphen version 2.2.2
## sift version sift5.2.2
## COSMIC version 78

## ESP version 20141103
## gencode version GENCODE 25
## genebuild version 2014-07
## HGMD-PUBLIC version 20162
## regbuild version 16
## assembly version GRCh38.p7
## ClinVar version 201610
## dbSNP version 147
## Column descriptions:
## Uploaded_variation : Identifier of uploaded variant
## Location : Location of variant in standard coordinate format (chr:start or chr:start-end)
## Allele : The variant allele used to calculate the consequence
## Gene : Stable ID of affected gene
## Feature : Stable ID of feature
## Feature_type : Type of feature - Transcript, RegulatoryFeature or MotifFeature
## Consequence : Consequence type
## cDNA_position : Relative position of base pair in cDNA sequence
## CDS_position : Relative position of base pair in coding sequence
## Protein_position : Relative position of amino acid in protein
## Amino_acids : Reference and variant amino acids
## Codons : Reference and variant codon sequence
## Existing_variation : Identifier(s) of co-located known variants
## Extra column keys:
## IMPACT : Subjective impact classification of consequence type
## DISTANCE : Shortest distance from variant to transcript
## STRAND : Strand of the feature (1/-1)
## FLAGS : Transcript quality flags
Tab-delimited output
The --tab flag instructs VEP to write output as a tab-delimited table.
This differs from the default output format in that each individual field from the "Extra" field is written to a separate tab-
delimited column.
This makes the output more suitable for import into spreadsheet programs such as Excel.
Furthermore the header is the same as the one for the VEP default output format and this is also the format used when selecting the
"TXT" option on the VEP web interface.
Example of VEP tab-delimited output format:
#Uploaded_variation Location Allele Gene Feature Feature_type
Consequence cDNA_position CDS_position Protein_position Amino_acids
Codons Existing_variation IMPACT DISTANCE STRAND FLAGS
11_224088_C/A 11:224088 A ENSG00000142082 ENST00000525319 Transcript
missense_variant 742 716 239 S/I
aGc/aTc - MODERATE - -1 -
11_224088_C/A 11:224088 A ENSG00000142082 ENST00000534381 Transcript
downstream_gene_variant - - - -
- - MODIFIER 1674 -1 -
11_224088_C/A 11:224088 A ENSG00000142082 ENST00000529055 Transcript
downstream_gene_variant - - - -
- - MODIFIER 134 -1 -
11_224585_G/A 11:224585 A ENSG00000142082 ENST00000529937 Transcript
intron_variant,NMD_transcript_variant - - - -
- - MODIFIER - -1 -
The choice and order of columns in the output may be configured using --fields. For instance:
./vep -i examples/homo_sapiens_GRCh38.vcf --cache --force_overwrite --tab --fields "Uploaded
variation,Location,Allele,Gene"

VCF output
The VEP script can also generate VCF output using the --vcf flag.
Main information about the specificity of the VEP VCF output format:
Consequences are added in the INFO field of the VCF file, using the key "CSQ" (you can change it using --vcf_info_field).
Data fields are encoded separated by the character "|" (pipe). The order of fields is written in the VCF header. Unpopulated fields
are represented by an empty string.
Output fields in the "CSQ" INFO field can be configured by using --fields.
Each prediction, for a given variant, is separated by the character "," in the CSQ INFO field (e.g. when a variant overlaps more than
1 transcript)
Here is a list of the (default) fields you can find within the CSQ field:
Allele|Consequence|IMPACT|SYMBOL|Gene|Feature_type|Feature|BIOTYPE|EXON|INTRON|HGVSc|HGVSp|cDNA_po
sition|CDS_position|Protein_position|Amino_acids|Codons|Existing_variation|DISTANCE|STRAND|FLAGS|S
YMBOL_SOURCE|HGNC_ID
Example of VEP command using the --vcf and --fields options:
./vep -i examples/homo_sapiens_GRCh38.vcf --cache --force_overwrite --vcf --fields
"Allele,Consequence,Feature_type,Feature"
VCFs produced by VEP can be filtered by filter_vep.pl in the same way as standard format output files.
If the input format was VCF, the file will remain unchanged save for the addition of the CSQ field and the header (unless using any
filtering). If an existing CSQ field is found, it will be replaced by the one added by the VEP (use --keep_csq to preserve it).
Custom data added with --custom are added as separate fields, using the key specified for each data file.
Commas in fields are replaced with ampersands (&) to preserve VCF format.
##INFO=<ID=CSQ,Number=.,Type=String,Description="Consequence annotations from Ensembl VEP. Format:
Allele|Consequence|IMPACT|SYMBOL|Gene|Feature_type|Feature|BIOTYPE|EXON|INTRON|HGVSc|HGVSp|cDNA_po
sition|CDS_position|Protein_position">
#CHROM POS ID REF ALT QUAL FILTER INFO
21 26978790 rs75377686 T C . .
CSQ=C|missense_variant|MODERATE|MRPL39|ENSG00000154719|Transcript|ENST00000419219|protein_coding|2
/8||ENST00000419219.1:c.251A>G|ENSP00000404426.1:p.Asn84Ser|260|251|84
JSON output
VEP can produce output in the form of serialised JSON objects using the --json flag. JSON is a serialisation format that can be parsed
and processed easily by many packages and programming languages; it is used as the default output format for Ensembl's REST
server .
Each input variant is reported as a single JSON object which constitutes one line of the output file. The JSON object is structured
somewhat differently to the other VEP output formats, in that per-variant fields (e.g. co-located existing variant details) are reported only
once. Consequences are grouped under the feature type that they affect (Transcript, Regulatory Feature, etc). The original input line
(e.g. from VCF input) is reported under the "input" key in order to aid aligning input with output. When using a cache file, frequencies for
co-located variants are reported by default (see --af_1kg, --af_gnomade).
Here follows an example of JSON output (prettified and redacted for display here):
{
"input": "1 1918090 test1 A G . . .",
"id": "test1",
"seq_region_name": "1",

"start": 1918090,
"end": 1918090,
"strand": 1,
"allele_string": "A/G",
"most_severe_consequence": "missense_variant",
"colocated_variants": [
{
"id": "COSV57068665",
"seq_region_name": "1",
"start": 1918090,
"end": 1918090,
"strand": 1,
"allele_string": "COSMIC_MUTATION"
},
{
"id": "rs28640257",
"seq_region_name": "1",
"start": 1918090,
"end": 1918090,
"strand": 1,
"allele_string": "A/G/T",
"minor_allele": "G",
"minor_allele_freq": 0.352,
"frequencies": {
"G": {
"amr": 0.5072,
"gnomade_sas": 0.369,
"gnomade": 0.4541,
"gnomade_oth": 0.4611,
"gnomade_asj": 0.3909,
"gnomade_nfe": 0.4944,
"gnomade_afr": 0.103,
"afr": 0.053,
"gnomade_amr": 0.5641,
"gnomade_fin": 0.474,
"sas": 0.3906,
"gnomade_eas": 0.4598,
"eur": 0.4901,
"eas": 0.4623
}
}
}
],
"transcript_consequences": [
{
"variant_allele": "G",
"consequence_terms": [
"missense_variant"
],
"gene_id": "ENSG00000178821",
"transcript_id": "ENST00000310991",
"strand": -1,
"cdna_start": 436,
"cdna_end": 436,
"cds_start": 422,
"cds_end": 422,
"protein_start": 141,
"protein_end": 141,
"codons": "aTg/aCg",
"amino_acids": "M/T",
"polyphen_prediction": "benign",
"polyphen_score": 0.001,
"sift_prediction": "tolerated",
"sift_score": 0.22,
"hgvsp": "ENSP00000311122.3:p.Met141Thr",
"hgvsc": "ENST00000310991.8:c.422T>C"
}

],
"regulatory_feature_consequences": [
{
"variant_allele": "G",
"consequence_terms": [
"regulatory_region_variant"
],
"regulatory_feature_id": "ENSR00000000255"
}
]
}
In accordance with JSON conventions, all keys (except alleles) are lower-case. Some keys also have different names and structures to
those found in the other VEP output formats:
Key JSON equivalent(s) Notes
Consequence consequence_terms
Gene gene_id
Feature transcript_id,
regulatory_feature_id,
motif_feature_id
Consequences are grouped under the feature type they affect
ALLELE variant_allele
SYMBOL gene_symbol
SYMBOL_SOURCE gene_symbol_source
ENSP protein_id
OverlapBP bp_overlap
OverlapPC percentage_overlap
Uploaded_variation id
Location seq_region_name,
start, end, strand
The variant's location field is broken down into constituent coordinate parts for clarity.
"seq_region_name" is used in place of "chr" or "chromosome" for consistency with other
parts of Ensembl's REST API
*_maf *_allele, *_maf
cDNA_position cdna_start, cdna_end
CDS_position cds_start, cds_end
Protein_position protein_start,
protein_end
SIFT sift_prediction,
sift_score
PolyPhen polyphen_prediction,
polyphen_score
Statistics
VEP writes an HTML file containing statistics pertaining to the results of your job; it is named [output_file]_summary.html (with the
default options the file will be named variant_effect_output.txt_summary.html). To view it, please open the file in your web browser.
To prevent VEP writing a stats file, use --no_stats.
To get a machine-readable text file in place of the HTML file, use --stats_text. You can get both a HTML file and plain text file by
using both --stats_text and --stats_html.
To change the name of the stats file from the default, use --stats_file [file].
The page contains several sections:
General statistics

General statistics
Summary of called consequence
types
Distribution of variants across
chromosomes
This section contains two tables. The first describes the cache and/or database used, the version of VEP, species, command line
parameters, input/output files and run time. The second table contains information about the number of variants, and the number of
genes, transcripts and regulatory features overlapped by the input.
Charts and tables
There then follows several charts, most with accompanying tables. Tables and charts are interactive; clicking on a row to highlight it in
the table will highlight the relevant segment in the chart, and vice versa.
!

./vep [options]
./vep --help
./vep --cache -i input.txt -o output.txt
Variant Effect Predictor Running VEP
VEP is run on the command line as follows (assuming you are in the ensembl-vep directory):
where [options] represent a set of flags and options. A basic set of flags can be listed using --help:
VEP can be run in the following modes:
For optimum performance, download a cache file for your species of interest, using either the installer or by following the VEP
Cache documentation, and run VEP with either the --cache or --offline option.
By connecting to the public Ensembl database servers in place of a cache. This can be adequate when annotating small files, but
the database servers can become busy and slow. To enable this option, use --database.
To run VEP using your own species and assembly, please use a --fasta file and --gff or --gtf annotation.
To run VEP with default options, use the following command:
where input.txt contains data in one of the compatible input formats and output.txt is the output file to be created.
Options can be passed as the full string (e.g. --format), or as the shortest unique string among the options (e.g. --form for --format, since
there is another option --force_overwrite).
You may use one or two hypen ("-") characters before each option name; --cache or -cache.
VEP options can also be read from:
Configuration files using --config. Options set in configuration files are overriden if specified on the command line.
Environment variables that start with prefix VEP_. For instance, you can set the cache flag with export VEP_CACHE=1 and the
input flag with export VEP_INPUT="/path/to/input.txt" before running ./vep. Options set in environment variables are
overriden if specified in configuration files or on the command line.
Basic options
Flag Alternat
e
Description Incompatib
le with
--help
! Display help message and quit !
--quiet
-q
Suppress warning messages.Not used by default --verbose
--verbose
-v
Print out a bit more information while running. Not used by default --quiet
--config [filename]
! Load configuration options from a config file. The config file should consist of
whitespace-separated pairs of option names and settings e.g.:
output_file my_output.txt
species mus_musculus
format vcf
host useastdb.ensembl.org
A config file can also be implicitly read; save the file as $HOME/.vep/vep.ini (or
!

equivalent directory if using --dir). Any options in this file will be overridden by
those specified in a config file using --config, and in turn by any options specified
on the command line. You can create a quick version file of this by setting the flags
as normal and running VEP in verbose (-v) mode. This will output lines that can be
copied to a config file that can be loaded in on the next run using --config. Not used
by default
--everything
-e
Shortcut flag to switch on all of the following:
--sift b, --polyphen b, --ccds, --hgvs, --symbol, --numbers, --domains, --regulatory, -
-canonical, --protein, --biotype, --af, --af_1kg, --af_esp, --af_gnomade, --
af_gnomadg, --max_af, --pubmed, --uniprot, --mane, --tsl, --appris, --variant_class,
--gene_phenotype, --mirna
!
--species [species]
! Species for your data. This can be the latin name e.g. "homo_sapiens" or any
Ensembl alias e.g. "mouse". Specifying the latin name can speed up initial
database connection as the registry does not have to load all available database
aliases on the server. Default = "homo_sapiens"
!
--assembly [name]
-a
Select the assembly version to use if more than one available. If using the cache,
you must have the appropriate assembly's cache file installed. If not specified and
you have only 1 assembly version installed, this will be chosen by default. Default
= use found assembly version
!
--input_file
[filename]
-i
Input file name. If not specified, VEP will attempt to read from STDIN. Can use
compressed file (gzipped).
!
--input_data
[string]
--id
Raw input data as a string. May be used, for example, to input a single rsID or
HGVS notation quickly to vep:
--input_data rs699
!
--format [format]
! Input file format - one of "ensembl", "vcf", "hgvs", "id", "region", "spdi".
By default, VEP auto-detects the input file format. Using this option you can specify
the input file is Ensembl, VCF, IDs, HGVS, SPDI or region format. Can use
compressed version (gzipped) of any file format listed above. Auto-detects format
by default
!
--output_file
[filename]
-o
Output file name. Results can write to STDOUT by specifying 'STDOUT' as the
output file name - this will force quiet mode. Default = "variant_effect_output.txt"
!
--force_overwrite
--force
By default, VEP will fail with an error if the output file already exists. You can force
the overwrite of the existing file by using this flag. Not used by default
!
--no_stats
! Don't generate a stats file. Provides marginal gains in run time. !
--stats_file
[filename]
--sf
Summary stats file name. This file contains a summary of the VEP run. If stats are
returned in an HTML file (default), the filename should end in .html or .htm.
Default = "variant_effect_output.txt_summary.html"
!
--stats_html
! Generate a HTML stats file (default). !
--stats_text
! Generate a plain text stats file. Can be combined with --stats_html to generate
both plain text and HTML stats files.
!
--warning_file
[filename]
! File name to write warnings and errors to. Default = STDERR (standard error) !
--
skipped_variants_fi
le [filename]
! File name to write skipped variants to. Default = STDERR (standard error) !
--max_sv_size
! Extend the maximum Structural Variant size VEP can process. !
--
no_check_variants_o
! Permit the use of unsorted input files. However running VEP on unsorted input files
slows down the tool and requires more memory.
!

rder
--fork [num_forks]
! Enable forking, using the specified number of forks. Forking can dramatically
improve runtime. Not used by default
!
Cache options
Flag Alternat
e
Description Output fields Incompatib
le with
--cache
! Enables use of the cache. Add --refseq or --merged to use the
refseq or merged cache, (if installed).
!
--database
--dir [directory]
! Specify the base cache/plugin directory to use. Default =
"$HOME/.vep/"
!
--dir_cache
[directory]
! Specify the cache directory to use. Default = "$HOME/.vep/" !
--dir_plugins
[directory]
! Specify the plugin directory to use. Default = "$HOME/.vep/" !
--offline
! Enable offline mode. No database connections will be made, and
a cache file or GFF/GTF file is required for annotation. Add --
refseq to use the refseq cache (if installed). Not used by default
!
--database
--check_svs
--fasta [file|dir]
--fa
Specify a FASTA file or a directory containing FASTA files to use to
look up reference sequence. The first time you run VEP with this
parameter an index will be built which can take a few minutes. This
is required if fetching HGVS annotations (--hgvs) or checking
reference sequences (--check_ref) in offline mode (--offline), and
optional with some performance increase in cache mode (--cache).
See documentation for more details. Not used by default
!
--refseq
! Specify this option if you have installed the RefSeq cache in order
for VEP to pick up the alternate cache directory. This cache
contains transcript objects corresponding to RefSeq transcripts.
Consequence output will be given relative to these transcripts in
place of the default Ensembl transcripts (see documentation)
REFSEQ_MA
TCH,
BAM_EDIT
--
gencode_b
asic
--merged
--merged
! Use the merged Ensembl and RefSeq cache. Consequences are
flagged with the SOURCE of each transcript used.
REFSEQ_MA
TCH,
BAM_EDIT,
SOURCE
--refseq
--cache_version
! Use a different cache version than the assumed default (the VEP
version). This should be used with Ensembl Genomes caches
since their version numbers do not match Ensembl versions. For
example, the VEP/Ensembl version may be 88 and the Ensembl
Genomes version 35. Not used by default
!
--show_cache_info
! Show source version information for selected cache and quit !
--buffer_size
[number]
! Sets the internal buffer size, corresponding to the number of
variants that are read in to memory simultaneously. Set this lower
to use less memory at the expense of longer run time, and higher
to use more memory with a faster run time. Default = 5000
!
Other annotation sources
Flag Alternate Description Output fields

--plugin [plugin
name]
! Use named plugin. Plugin modules should be installed in the Plugins
subdirectory of the VEP cache directory (defaults to $HOME/.vep/).
Multiple plugins can be used by supplying the --plugin flag multiple times.
See plugin documentation. Not used by default
Plugin-dependent
--custom file=
[filename]
! Add custom annotation to the output. Files must be tabix indexed or in
the bigWig format. Multiple files can be specified by supplying the --
custom flag multiple times. See here for full details. Not used by default
SOURCE, Custom
file dependent
--gff [filename]
! Use GFF transcript annotations in [filename] as an annotation source.
Requires a FASTA file of genomic sequence. Not used by default
SOURCE
--gtf [filename]
! Use GTF transcript annotations in [filename] as an annotation source.
Requires a FASTA file of genomic sequence. Not used by default
SOURCE
--bam [filename]
! ADVANCED Use BAM file of sequence alignments to correct transcript
models not derived from reference genome sequence. Used to correct
RefSeq transcript models. Enables --use_transcript_ref; add --
use_given_ref to override this behaviour. Not used by default
BAM_EDIT
--
use_transcript_ref
! By default VEP uses the reference allele provided in the input file to
calculate consequences for the provided alternate allele(s). Use this flag
to force VEP to replace the provided reference allele with sequence
derived from the overlapped transcript. This is especially relevant when
using the RefSeq cache, see documentation for more details. The
GIVEN_REF and USED_REF fields are set in the output to indicate any
change. Not used by default
GIVEN_REF,
USED_REF
--use_given_ref
! Using --bam or a BAM-edited RefSeq cache by default enables --
use_transcript_ref; add this flag to override this behaviour and use the
provided reference allele from the input. Not used by default
!
--
custom_multi_alleli
c
! By default, comma separated lists found within the INFO field of custom
annotation VCFs are assumed to be allele specific. For example, a
variant with allele_string A/G/C with associated custom annotation
'single,double,triple' will associate triple with C, double with G and single
with A. This flag instructs VEP to return all annotations for all alleles. Not
used by default
!
Output format options
Flag Alternate Description Output
fields
Incompatib
le with
--vcf
! Writes output in VCF format. Consequences are added in the
INFO field of the VCF file, using the key "CSQ". Data fields are
encoded separated by "|"; the order of fields is written in the VCF
header. Output fields in the "CSQ" INFO field can be selected by
using --fields.
If the input format was VCF, the file will remain unchanged save for
the addition of the CSQ field (unless using any filtering).
Custom data added with --custom are added as separate fields,
using the key specified for each data file.
Commas in fields are replaced with ampersands (&) to preserve
VCF format.
Not used by default
!
--json
--tab
--summary
--
most_sever
e
--tab
! Writes output in tab-delimited format. Not used by default !
--json
--vcf
--json
! Writes output in JSON format. Not used by default !
--tab
--vcf
--compress_output
! Writes output compressed using either gzip or bgzip. Not used by !

[gzip|bgzip]
default
--fields [list]
! Configure the output format using a comma separated list of fields.
Can only be used with tab (--tab) or VCF format (--vcf) output.
For the tab format output, the selected fields may be those present
in the default output columns, or any of those that appear in the
Extra column (including those added by plugins or custom
annotations) if the appropriate output is available (e.g. use --
show_ref_allele to access 'REF_ALLELE'). Output remains tab-
delimited.
For the VCF format output, the selected fields are those present
within the "CSQ" INFO field.
Example of command for the tab output:
--tab --fields
"Uploaded_variation,Location,Allele,Gene"
Example of command for the VCF format output:
--vcf --fields
"Allele,Consequence,Feature_type,Feature"
Not used by default
!
--minimal
! Convert alleles to their most minimal representation before
consequence calculation i.e. sequence that is identical between
each pair of reference and alternate alleles is trimmed off from
both ends, with coordinates adjusted accordingly.
Note this may lead to discrepancies between input coordinates
and coordinates reported by VEP relative to transcript sequences;
to avoid issues, use --allele_number and/or ensure that your input
variants have unique identifiers. The MINIMISED flag is set in the
VEP output where relevant. For an insertion/deletion, the allele is
minimised by default. To access the input allele before
minimisation, use --uploaded_allele.
Not used by default
MINIMISED
--individual
Output options
Flag Alternat
e
Description Output
fields
Incompatib
le with
--variant_class
! Output the Sequence Ontology variant class. Not used by default VARIANT_
CLASS
!
--sift [p|s|b]
! Species limited SIFT predicts whether an amino acid substitution
affects protein function based on sequence homology and the
physical properties of amino acids. VEP can output the prediction
term, score or both. Not used by default
SIFT
--
most_sever
e
--summary
--polyphen [p|s|b]
! Human only PolyPhen is a tool which predicts possible impact of
an amino acid substitution on the structure and function of a human
protein using straightforward physical and comparative
considerations. VEP can output the prediction term, score or both.
VEP uses the humVar score by default - use --humdiv to retrieve the
humDiv score. Not used by default
PolyPhen
--
most_sever
e
--summary
--humdiv
! Human only Retrieve the humDiv PolyPhen prediction instead of
the default humVar. Not used by default
PolyPhen !
--nearest
[transcript|gene|sy
mbol]
! Retrieve the transcript or gene with the nearest protein-coding
transcription start site (TSS) to each input variant. Use "transcript" to
retrieve the transcript stable ID, "gene" to retrieve the gene stable ID,
NEAREST !

or "symbol" to retrieve the gene symbol. Note that the nearest TSS
may not belong to a transcript that overlaps the input variant, and
more than one may be reported in the case where two are
equidistant from the input coordinates.
Currently only available when using a cache annotation source, and
requires the Set::IntervalTree perl module.
Not used by default
--distance
[bp_distance(,downs
tream_distance)]
! Modify the distance up and/or downstream between a variant and a
transcript for which VEP will assign the upstream_gene_variant or
downstream_gene_variant consequences. Giving one distance will
modify both up- and downstream distances; prodiving two separated
by commas will set the up- (5') and down- (3') stream distances
respectively. Default: 5000
!
--overlaps
! Report the proportion and length of a transcript overlapped by a
structural variant in VCF format.
!
--gene_phenotype
! Indicates if the overlapped gene is associated with a phenotype,
disease or trait. See list of phenotype sources. Not used by default
GENE_PH
ENO
!
--regulatory
! Look for overlaps with regulatory regions. VEP can also report if a
variant falls in a high information position within a transcription factor
binding site. Output lines have a Feature type of RegulatoryFeature
or MotifFeature. Not used by default
MOTIF_NA
ME,
MOTIF_PO
S,
HIGH_INF_
POS,
MOTIF_SC
ORE_CHA
NGE
!
--cell_type
! Report only regulatory regions that are found in the given cell type(s).
Can be a single cell type or a comma-separated list. The functional
type in each cell type is reported under CELL_TYPE in the output. To
retrieve a list of cell types, use --cell_type list. Not used by default
CELL_TYP
E
!
--individual
[all|ind list]
! Consider only alternate alleles present in the genotypes of the
specified individual(s). May be a single individual, a comma-
separated list or "all" to assess all individuals separately. Individual
variant combinations homozygous for the given reference allele will
not be reported. Each individual and variant combination is given on
a separate line of output. Only works with VCF files containing
individual genotype data; individual IDs are taken from column
headers. Not used by default
IND, ZYG
--minimal
--
individual_z
yg
--individual_zyg
[all|ind list]
! Consider alternate and reference alleles present in the genotypes of
the specified individual(s). May be a single individual, a comma-
separated list or "all" to assess all individuals separately. Returns a
list of individuals and their zygosity. Only works with VCF files
containing individual genotype data; individual IDs are taken from
column headers. Not used by default
ZYG
--individual
--phased
! Force VCF genotypes to be interpreted as phased. For use with
plugins that depend on phased data. Not used by default
!
--allele_number
! Identify allele number from VCF input, where 1 = first ALT allele, 2 =
second ALT allele etc. Useful when using --minimal Not used by
default
ALLELE_N
UM
!
--show_ref_allele
! Adds the reference allele in the output (after minimisation). Mainly
useful for the VEP "default" and tab-delimited output formats. Not
used by default
REF_ALLE
LE
!
--uploaded_allele
! Adds the uploaded allele string in the output (before minimisation). UPLOADE
D_ALLELE
!
--total_length
! Give cDNA, CDS and protein positions as Position/Length. Not used
by default
!
--numbers
! Adds affected exon and intron numbering to to output. Format is
Number/Total. Not used by default
EXON,
INTRON
--
most_sever
e

--summary
--mirna
! Reports where the variant lies in the miRNA secondary structure. Not
used by default
! !
--no_escape
! Don't URI escape HGVS strings. Default = escape !
--keep_csq
! Don't overwrite existing CSQ entry in VCF INFO field. Overwrites by
default
!
--vcf_info_field
[CSQ|ANN|(other)]
! Change the name of the INFO key that VEP write the consequences
to in its VCF output. Use "ANN" for compatibility with other tools such
as snpEff . Default: CSQ
!
--terms
[SO|display|NCBI]
-t
The type of consequence terms to output. The Ensembl terms are
described here. The Sequence Ontology is a joint effort by genome
annotation centres to standardise descriptions of biological
sequences. Default = "SO"
!
--no_headers
! Don't write header lines in output files. Default = add headers !
--shift_3prime
[0|1]
! Right aligns all variants relative to their associated transcripts prior to
consequence calculation.
An example using this option can be found here.
Default = 0
--shift_hgvs
--shift_genomic
[0|1]
! Right aligns all variants, including intergenic variants, before
consequence calculation and updates the Location field.
An example using this option can be found here.
Default = 0
--shift_hgvs
--shift_length
! Reports the distance each variant has been shifted when used in
conjuction with --shift_3prime
Identifiers
Flag Alternat
e
Description Output
fields
Incompatib
le with
--hgvs
! Add HGVS nomenclature based on Ensembl stable identifiers to
the output. Both coding and protein sequence names are added
where appropriate. To generate HGVS identifiers when using --cache
or --offline you must use a FASTA file and --fasta. HGVS notations
given on Ensembl identifiers are versioned. Not used by default
HGVSc,
HGVSp,
HGVS_OFF
SET
!
--hgvsg
! Add genomic HGVS nomenclature based on the input
chromosome name. To generate HGVS identifiers when using --
cache or --offline you must use a FASTA file and --fasta. Not used by
default
HGVSg !
--
hgvsg_use_accession
! Force --hgvsg to return RefSeq reference sequence. For example,
reports NC_000002.11 for human chromosome 2 (build GRCh38).
HGVSg !
--
hgvsp_use_predictio
n
! Force --hgvs to return the HGVSp notation in predicted format. For
example, ENSP00000233741.4:p.Thr367AsnfsTer13 will be returned
as ENSP00000233741.4:p.(Thr367AsnfsTer13).
HGVSp !
--ambiguous_hgvs
[0|1]
! Allow input HGVSp to resolve to all genomic locations. Otherwise,
most likely transcript will be selected. Default: 0 (most likely transcript
selected)
! !
--spdi
! Add genomic SPDI notation. To generate SPDI when using --cache
or --offline you must use a FASTA file and --fasta. Not used by default
SPDI !
--ga4gh_vrs
! Add GA4GH Variation Representation Specification (VRS) notation.
To generate GA4GH VRS when using --cache or --offline you must
use a FASTA file and --fasta. Not used by default
GA4GH_V
RS
--vcf

--shift_hgvs [0|1]
! Enable or disable 3' shifting of HGVS notations. HGVS nomenclature
requires an ambiguous sequence change to be described at the most
3' possible location. When enabled, this causes "shifting" to the most
3' possible coordinates (relative to the transcript sequence and
strand) before the HGVS notations are calculated; the flag
HGVS_OFFSET is set to the number of bases by which the variant
has shifted, relative to the input genomic coordinates. If
HGVS_OFFSET is equals to 0, no value will be added to
HGVS_OFFSET column. To disable the changing of location at
transcript level set --shift_hgvs to 0. Default: 1 (shift)
!
--
transcript_version
! Add version numbers to Ensembl transcript identifiers !
--protein
! Add the Ensembl protein identifier to the output where appropriate.
Not used by default
ENSP
--
most_sever
e
--summary
--symbol
! Adds the gene symbol (e.g. HGNC) (where available) to the output.
Some gene symbol, e.g. HGNC, are only available in merged cache
and therefore should be used with --merged option while using cache
to get result. Not used by default
SYMBOL,
SYMBOL_S
OURCE,
HGNC_ID
--
most_sever
e
--summary
--ccds
! Adds the CCDS transcript identifer (where available) to the output.
Not used by default
CCDS
--
most_sever
e
--summary
--uniprot
! Adds best match accessions for translated protein products from
three UniProt -related databases (SWISSPROT, TREMBL and
UniParc) to the output. Not used by default
SWISSPRO
T, TREMBL,
UNIPARC,
UNIPROT_I
SOFORM
--
most_sever
e
--summary
--tsl
! Adds the transcript support level for this transcript to the output. Not
used by default
TSL
--
most_sever
e
--summary
--appris
! Adds the APPRIS isoform annotation for this transcript to the output.
Not used by default
APPRIS
--
most_sever
e
--summary
--canonical
! Adds a flag indicating if the transcript is the canonical transcript for
the gene. Not used by default
CANONICA
L
--
most_sever
e
--summary
--mane
! Adds a flag indicating if the transcript is the MANE Select or MANE
Plus Clinical transcript for the gene. Not used by default
MANE_SEL
ECT,
MANE_PLU
S_CLINICA
L
--
most_sever
e
--summary
--mane_select
! Adds a flag indicating if the transcript is the MANE Select transcript
for the gene. Not used by default
MANE_SEL
ECT
--
most_sever
e
--summary
--biotype
! Adds the biotype of the transcript or regulatory feature. Not used by
default
BIOTYPE
--
most_sever
e
--summary
--domains
! Adds names of overlapping protein domains to output. Not used by
default
DOMAINS
--
most_sever

e
--summary
--xref_refseq
! Output aligned RefSeq mRNA identifier for transcript. Not used by
default
RefSeq
--
most_sever
e
--summary
--synonyms [file]
! Load a file of chromosome synonyms. File should be tab-delimited
with the primary identifier in column 1 and the synonym in column 2.
Synonyms allow different chromosome identifiers to be used in the
input file and any annotation source (cache, database, GFF, custom
file, FASTA file). Not used by default
! !
Co-located variants
Flag Alternat
e
Description Output
fields
Incompatib
le with
--check_existing
! Checks for the existence of known variants that are co-located with
your input. By default the alleles are compared and variants on an
allele-specific basis - to compare only coordinates, use --
no_check_alleles.
Some databases may contain variants with unknown (null) alleles
and these are included by default; to exclude them use --
exclude_null_alleles.
See this page for more details.
Not used by default
Existing_va
riation,
CLIN_SIG,
SOMATIC,
PHENO
!
--check_svs
! Checks for the existence of structural variants that overlap your input.
Currently requires database access. Not used by default
SV --offline
--clin_sig_allele
[1|0]
! Return allele specific clinical significance. Setting this option to 0 will
provide all known clinical significance values at the given locus.
Default: 1 (Provide allele-specific annotations)
CLIN_SIG !
--
exclude_null_allele
s
! Do not include variants with unknown alleles when checking for co-
located variants. Our human database contains variants from HGMD
and COSMIC for which the alleles are not publically available; by
default these are included when using --check_existing, use this flag
to exclude them. Not used by default
!
--no_check_alleles
! When checking for existing variants, by default VEP only reports a
co-located variant if none of the input alleles are novel. For example,
if your input variant has alleles A/G, and an existing co-located
variant has alleles A/C, the co-located variant will not be reported.
Strand is also taken into account - in the same example, if the input
variant has alleles T/G but on the negative strand, then the co-
located variant will be reported since its alleles match the reverse
complement of input variant.
Use this flag to disable this behaviour and compare using
coordinates alone. Not used by default
!
--af
! Add the global allele frequency (AF) from 1000 Genomes Phase 3
data for any known co-located variant to the output. For this and all --
af_* flags, the frequency reported is for the input allele only, not
necessarily the non-reference or derived allele. Not used by default
AF !
--max_af
! Report the highest allele frequency observed in any population from
1000 genomes, ESP or gnomAD. Not used by default
MAX_AF,
MAX_AF_P
OPS
--database

--af_1kg
! Add allele frequency from continental populations
(AFR,AMR,EAS,EUR,SAS) of 1000 Genomes Phase 3 to the
output. Must be used with --cache. Not used by default
AFR_AF,
AMR_AF,
EAS_AF,
EUR_AF,
SAS_AF
--database
--af_esp
! Include allele frequency from NHLBI-ESP populations. Must be
used with --cache. Deprecated.
AA_AF,
EA_AF
--database
--af_gnomade
--
af_gnom
ad
Include allele frequency from Genome Aggregation Database
(gnomAD) exome populations. Note only data from the gnomAD
exomes are included; to retrieve data from the additional genomes
data set, see this guide. Must be used with --cache Not used by
default
gnomADe_
AF,
gnomADe_
AFR_AF,
gnomADe_
AMR_AF,
gnomADe_
ASJ_AF,
gnomADe_
EAS_AF,
gnomADe_
FIN_AF,
gnomADe_
NFE_AF,
gnomADe_
OTH_AF,
gnomADe_
SAS_AF
--database
--af_gnomadg
! Include allele frequency from Genome Aggregation Database
(gnomAD) genome populations. Note only data from the gnomAD
genomes are included; to retrieve data from the additional genomes
data set, see this guide. Must be used with --cache Not used by
default
gnomADg_
AF,
gnomADg_
AFR_AF,
gnomADg_
AMI_AF,
gnomADg_
AMR_AF,
gnomADg_
ASJ_AF,
gnomADg_
EAS_AF,
gnomADg_
FIN_AF,
gnomADg_
MID_AF,
gnomADg_
NFE_AF,
gnomADg_
OTH_AF,
gnomADg_
SAS_AF
--database
--af_exac
! Include allele frequency from ExAC project populations. Must be
used with --cache. Deprecated.
ExAC_AF,
ExAC_Adj_
AF,
ExAC_AFR
_AF,
ExAC_AMR
_AF,
ExAC_EAS
_AF,
ExAC_FIN_
AF,
ExAC_NFE
_AF,
ExAC_OTH
_AF,
ExAC_SAS
_AF
--database
--pubmed
! Report Pubmed IDs for publications that cite existing variant. Must be PUBMED
--database

used with --cache. Not used by default
--var_synonyms
! Report known synonyms for co-located variants. Must be used with --
cache. Not used by default
VAR_SYNO
NYMS
--database
--failed [0|1]
! When checking for co-located variants, by default VEP will exclude
variants that have been flagged as failed. Set this flag to include such
variants. Default: 0 (exclude)
!
Filtering and QC options
NOTE: The filtering options here filter your results before they are written to your output file. Using VEP's filtering script, it is possible to
filter your results after VEP has run. This way you can retain all of the results and run multiple filter sets on the same results to find
different data of interest.
Flag Alternat
e
Description Output
fields
Incompatib
le with
--gencode_basic
! Limit your analysis to transcripts belonging to the GENCODE basic
set. This set has fragmented or problematic transcripts removed. Not
used by default
!
--refseq
--exclude_predicted
! When using the RefSeq or merged cache, exclude predicted
transcripts (i.e. those with identifiers beginning with "XM_" or "XR_").
!
--transcript_filter
! ADVANCED Filter transcripts according to any arbitrary set of rules.
Uses similar notation to filter_vep.
You may filter on any key defined in the root of the transcript object;
most commonly this will be "stable_id":
--transcript_filter "stable_id match N[MR]_"
!
--check_ref
! Force VEP to check the supplied reference allele against the
sequence stored in the Ensembl Core database or supplied FASTA
file. Lines that do not match are skipped. Checking is done on the
minimised sequence. Example chr13 32900399 . AGT A . the As are
removed and the reference sequence is checked from 32900400 to
see if it matches GTNot used by default
!
--lookup_ref
--lookup_ref
! Force overwrite the supplied reference allele with the sequence
stored in the Ensembl Core database or supplied FASTA file. Not
used by default
!
--check_ref
--dont_skip
! Don't skip input variants that fail validation, e.g. those that fall on
unrecognised sequences.
Combining --check_ref with --dont_skip will add a CHECK_REF
output field when the given reference does not match the underlying
reference sequence.
CHECK_REF
--allow_non_variant
! When using VCF format as input and output, by default VEP will skip
non-variant lines of input (where the ALT allele is null). Enabling this
option the lines will be printed in the VCF output with no
consequence data added.
!
--chr [list]
! Select a subset of chromosomes to analyse from your file. Any data
not on this chromosome in the input will be skipped. The list can be
comma separated, with "-" characters representing an interval.
For example, to include chromosomes 1, 2, 3, 10 and X you could
use --chr 1-3,10,X Not used by default
!
--coding_only
! Only return consequences that fall in the coding regions of
transcripts. Not used by default
!
--
most_sever
e
--summary
--no_intergenic
! Do not include intergenic consequences in the output. Not used by !
--

default most_sever
e
--summary
--pick
! Pick one line or block of consequence data per variant, including
transcript-specific columns.
Consequences are chosen according to the criteria described here,
and the order the criteria are applied may be customised with --
pick_order. This is the best method to use if you are interested only
in one consequence per variant. Not used by default
!
--
most_sever
e
--summary
--pick_allele
! Like --pick, but chooses one line or block of consequence data per
variant allele. Will only differ in behaviour from --pick when the input
variant has multiple alternate alleles. Not used by default
!
--
most_sever
e
--summary
--per_gene
! Output only the most severe consequence per gene. The transcript
selected is arbitrary if more than one has the same predicted
consequence. Uses the same ranking system as --pick. Not used by
default
!
--pick_allele_gene
! Like --pick_allele, but chooses one line or block of consequence data
per variant allele and gene combination. Not used by default
!
--flag_pick
! As per --pick, but adds the PICK flag to the chosen block of
consequence data and retains others. Not used by default
PICK
--
most_sever
e
--summary
--flag_pick_allele
! As per --pick_allele, but adds the PICK flag to the chosen block of
consequence data and retains others. Not used by default
PICK
--
most_sever
e
--summary
--
flag_pick_allele_ge
ne
! As per --pick_allele_gene, but adds the PICK flag to the chosen block
of consequence data and retains others. Not used by default
PICK !
--pick_order
[c1,c2,...,cN]
! Customise the order of criteria (and the list of criteria) applied when
choosing a block of annotation data with one of the following options:
--pick, --pick_allele, --per_gene, --pick_allele_gene, --flag_pick, --
flag_pick_allele, --flag_pick_allele_gene. See this page for the default
order.
Valid criteria are: mane_select, mane_plus_clinical, canonical, appris,
tsl, biotype, ccds, rank, length, ensembl, refseq. e.g.:
--pick --pick_order tsl,appris,rank
!
--most_severe
! Output only the most severe consequence per variant. Transcript-
specific columns will be left blank. Consequence ranks are given in
this table.
To include regulatory consequences, use the --regulatory option in
combination with this flag.
Not used by default
!
--appris
--biotype
--canonical
--ccds
--
coding_only
--domains
--flag_pick
--
flag_pick_al
lele
--
no_intergen
ic
--numbers

--pick
--pick_allele
--polyphen
--protein
--sift
--summary
--symbol
--tsl
--uniprot
--
xref_refseq
--summary
! Output only a comma-separated list of all observed consequences
per variant. Transcript-specific columns will be left blank. Not used by
default
!
--appris
--biotype
--canonical
--ccds
--
coding_only
--domains
--flag_pick
--
flag_pick_al
lele
--
most_sever
e
--
no_intergen
ic
--numbers
--pick
--pick_allele
--polyphen
--protein
--sift
--symbol
--tsl
--uniprot
--
xref_refseq
--filter_common
! Shortcut flag for the filters below - this will exclude variants that have
a co-located existing variant with global AF > 0.01 (1%). May be
modified using any of the following freq_* filters. Not used by default
FREQS !
--check_frequency
! Turns on frequency filtering. Use this to include or exclude variants
based on the frequency of co-located existing variants in the
Ensembl Variation database. You must also specify all of the --freq_*
flags below. Frequencies used in filtering are added to the output
under the FREQS key in the Extra field. Not used by default
FREQS !
--freq_pop [pop]
! Name of the population to use in frequency filter. This must be one of
the following:
Name Description
1KG_ALL 1000 genomes combined population (global)
1KG_AFR 1000 genomes combined African population
1KG_AMR 1000 genomes combined American population
1KG_EAS 1000 genomes combined East Asian population
!

1KG_EUR 1000 genomes combined European population
1KG_SAS 1000 genomes combined South Asian population
gnomADe gnomAD exomes combined population
gnomADe_AFR gnomAD exomes African/African American
population
gnomADe_AMR gnomAD exomes Latino population
gnomADe_ASJ gnomAD exomes Ashkenazi Jewish population
gnomADe_EAS gnomAD exomes East Asian population
gnomADe_FIN gnomAD exomes Finnish population
gnomADe_NFE gnomAD exomes non-Finnish European
population
gnomADe_OTH gnomAD exomes other population
gnomADe_SAS gnomAD exomes South Asian population
gnomADg gnomAD genomes combined population
gnomADg_AFR gnomAD genomes African/African American
population
gnomADg_AMR gnomAD genomes Latino population
gnomADg_AMI gnomAD genomes Amish population
gnomADg_ASJ gnomAD genomes Ashkenazi Jewish population
gnomADg_EAS gnomAD genomes East Asian population
gnomADg_FIN gnomAD genomes Finnish population
gnomADg_MID gnomAD genomes Mid-eastern population
gnomADg_NFE gnomAD genomes non-Finnish European
population
gnomADg_OTH gnomAD genomes other population
gnomADg_SAS gnomAD genomes South Asian population
--freq_freq [freq]
! Allele frequency to use for filtering. Must be a float value between 0
and 1
!
--freq_gt_lt
[gt|lt]
! Specify whether the frequency of the co-located variant must be
greater than (gt) or less than (lt) the value specified with --freq_freq
!
--freq_filter
[exclude|include]
! Specify whether to exclude or include only variants that pass the
frequency filter
!
Database options
Flag Alternate Description Output
fields
Incompatible
with
--database
! Enable VEP to use local or remote databases. !
--af_1kg
--af_esp
--af_exac
--af_gnomade
--af_gnomadg
--cache
--max_af
--offline

--pubmed
--
var_synonyms
--host [hostname]
! Manually define the database host to connect to. Users in the US
may find connection and transfer speeds quicker using our East
coast mirror, useastdb.ensembl.org. Default =
"ensembldb.ensembl.org"
!
--user [username]
-u
Manually define the database username. Default = "anonymous" !
--password
[password]
--pass
Manually define the database password. Not used by default !
--port [number]
! Manually define the database port. Default = 5306 !
--genomes
! Override the default connection settings with those for the
Ensembl Genomes public MySQL server. Required when using
any of the Ensembl Genomes species. Not used by default
!
--is_multispecies
[0|1]
! Some of the Ensembl Genomes databases (mainly bacteria and
protists) are composed of a collection of close species. It updates
the database connection settings (i.e. the database name) if the
value is set to 1. Default: 0
!
--lrg
! Map input variants to LRG coordinates (or to chromosome
coordinates if given in LRG coordinates), and provide
consequences on both LRG and chromosomal transcripts. Not
used by default
!
--offline
--db_version
[number]
! Force VEP to connect to a specific version of the Ensembl
databases. Not recommended as there may be conflicts between
software and database versions. Not used by default
!
--registry
[filename]
! Defining a registry file overwrites other connection settings and
uses those found in the specified registry file to connect. Not used
by default
!
!

Variant Effect Predictor Annotation sources
VEP can use a variety of annotation sources to retrieve the transcript models used to predict consequence types.
Cache - a downloadable file containing all transcript models, regulatory features and variant data for a species
GFF or GTF - use transcript models defined in a tabix-indexed GFF or GTF file
Requires a FASTA file in --offline mode or if the desired species or assembly is not part of the Ensembl species list.
Database - connect to a MySQL database server hosting Ensembl databases
Data from VCF, BED and bigWig files can also be incorporated by VEP's Custom annotation feature.
Using a cache is the most efficient way to use VEP; we would encourage you to use a cache wherever
possible. Caches are easy to download and set up using the installer. Follow the tutorial for a simple
guide.
Caches
Using a cache (--cache) is the fastest and most efficient way to use VEP, as in most cases only a single initial network connection is
made and most data is read from local disk. Use offline mode to eliminate all network connections for speed and/or privacy.
Downloading caches
Ensembl creates cache files for every species for each Ensembl release. They can be automatically downloaded and configured using
INSTALL.pl.
If interested in RefSeq transcripts you may download an alternate cache file (e.g. homo_sapiens_refseq), or a merged file of RefSeq and
Ensembl transcripts (eg homo_sapiens_merged); remember to specify --refseq or --merged when running VEP to use the relevant
cache. See documentation for full details.
Manually downloading caches
It is also simple to download and set up caches without using the installer. By default, VEP searches for caches in $HOME/.vep; to use a
different directory when running VEP, use --dir_cache.
Indexed cache (https://ftp.ensembl.org/pub/release-112/variation/indexed_vep_cache/)
Essential for human and other species with large sets of variant data - requires Bio::DB::HTS (setup by INSTALL.pl) or tabix ,
e.g.:
cd $HOME/.vep
curl -O
https://ftp.ensembl.org/pub/release-112/variation/indexed_vep_cache/homo_sapiens_vep_112_GRCh
38.tar.gz
tar xzf homo_sapiens_vep_112_GRCh38.tar.gz
Non-indexed cache (https://ftp.ensembl.org/pub/release-112/variation/vep/), e.g.:
cd $HOME/.vep
curl -O
https://ftp.ensembl.org/pub/release-112/variation/vep/homo_sapiens_vep_112_GRCh38.tar.gz
tar xzf homo_sapiens_vep_112_GRCh38.tar.gz
FTP directories by species grouping:
Ensembl: Vertebrates (indexed)
Ensembl Genomes: Bacteria | Fungi (indexed) | Metazoa (indexed) | Plants (indexed) | Protists (indexed)

NB: When using Ensembl Genomes caches, you should use the --cache_version option to specify the relevant Ensembl Genomes
version number as these differ from the concurrent Ensembl/VEP version numbers.
Data in the cache
The data content of VEP caches vary by species. This table shows the contents of the default human cache files in release 112.
Source Version (GRCh38) Version (GRCh37)
Ensembl database version 112 112
Genome assembly GRCh38.p14 GRCh37.p13
MANE Version v1.3 n/a
GENCODE 46 19
RefSeq GCF_000001405.40-RS_2023_10
(GCF_000001405.40_GRCh38.p14_genomic.gff)
105.20220307
(GCF_000001405.25_GRCh37.p13_genomic.gff)
Regulatory build 1.0 1.0
PolyPhen 2.2.3 2.2.2
SIFT 6.2.1 5.2.2
dbSNP 156 156
COSMIC 98 98
HGMD-PUBLIC 2020.4 2020.4
ClinVar 2023-10 2023-06
1000 Genomes Phase 3 (remapped) Phase 3
gnomAD exomes r2.1.1, exomes only r2.1, exomes only
gnomAD genomes r3.1.2, genomes only !
Convert with tabix
If you have Bio::DB::HTS (as set up by INSTALL.pl) or tabix installed on your system, the speed of retrieving existing co-located
variants can be greatly improved by converting the cache files using the supplied script, convert_cache.pl. This replaces the plain-text,
chunked variant dumps with a single tabix-indexed file per chromosome. The script is simple to run:
perl convert_cache.pl -species [species] -version [vep_version]
To convert all species and all versions, use "all":
perl convert_cache.pl -species all -version all
A full description of the options can be seen using --help. When complete, VEP will automatically detect the converted cache and use
this in place.
Note that tabix and bgzip must be installed on your system to convert a cache. INSTALL.pl downloads these when setting up
Bio::DB::HTS; to enable convert_cache.pl to find them, run:
export PATH=${PATH}:${PWD}/htslib
Data privacy and offline mode
When using the public database servers, VEP requests transcript and variation data that overlap the loci in your input file. As such, these
coordinates are transmitted over the network to a public server, which may not be appropriate for the analysis of sensitive or private data.
To run VEP in an offline mode that does not use any network connections, use the flag --offline.
The limitations described above apply absolutely when using offline mode. For example, if you specify --offline and --format id, VEP will
report an error and refuse to run:

ERROR: Cannot use ID format in offline mode
All other features, including the ability to use custom annotations and plugins, are accessible in offline mode.
GFF/GTF files
VEP can use transcript annotations defined in GFF or GTF files. The files must be bgzipped and indexed with tabix and a FASTA file
containing the genomic sequence is required in order to generate transcript models. This allows you to run VEP on data from any
species and assembly.
Your GFF or GTF file must be sorted in chromosomal order. VEP does not use header lines so it is safe to remove them.
grep -v "#" data.gff | sort -k1,1 -k4,4n -k5,5n -t$'\t' | bgzip -c > data.gff.gz
tabix -p gff data.gff.gz
./vep -i input.vcf --gff data.gff.gz --fasta genome.fa.gz
You may use any number of GFF/GTF files in this way, providing they refer to the same genome. You may also use them in concert with
annotations from a cache or database source; annotations are distinguished by the SOURCE field in the VEP output.
GFF file
Example of command line with GFF, using flag --gff :
./vep -i input.vcf --cache --gff data.gff.gz --fasta genome.fa.gz
NOTE: If you wish to customise the name of the GFF as it appears in the SOURCE field and VEP output header, use the longer --
custom annotation form:
--custom file=data.gff.gz,short_name=frequency,format=gff
GTF file
Example of command line with GTF, using flag --gtf :
./vep -i input.vcf --cache --gtf data.gtf.gz --fasta genome.fa.gz
NOTE: If you wish to customise the name of the GFF as it appears in the SOURCE field and VEP output header, use the longer --
custom annotation form:
--custom file=data.gtf.gz,short_name=frequency,format=gtf
GFF format expectations
VEP has been tested on GFF files generated by Ensembl and NCBI (RefSeq). Due to inconsistency in the GFF specification and
adherence to it, VEP may encounter problems parsing some GFF files. For the same reason, not all transcript biotypes defined in your
GFF may be supported by VEP. VEP does not support GFF files with embedded FASTA sequence.
Column "type" (3rd column):
The following entity/feature types are supported by VEP. Lines of other types will be ignored; if this leads to an incomplete transcript
model, the whole transcript model may be discarded.
Show supported types
Expected parameters in the 9th column:

ID
Only required for the genes and transcripts entities.
parent/Parent
- Entities in the GFF are expected to be linked using a key named "parent" or "Parent" in the attributes (9th) column of the GFF.
- Unlinked entities (i.e. those with no parents or children) are discarded.
- Sibling entities (those that share the same parent) may have overlapping coordinates, e.g. for exon and CDS entities.
biotype
Transcripts require a Sequence Ontology biotype to be defined in order to be parsed by VEP.
The simplest way to define this is using an attribute named "biotype" on the transcript entity. Other configurations are supported in
order for VEP to be able to parse GFF files from NCBI and other sources.
Here is an example:
##gff-version 3.2.1
##sequence-region 1 1 10000
1 Ensembl gene 1000 5000 . + . ID=gene1;Name=GENE1
1 Ensembl transcript 1100 4900 . + .
ID=transcript1;Name=GENE1-001;Parent=gene1;biotype=protein_coding
1 Ensembl exon 1200 1300 . + . ID=exon1;Name=GENE1-001_1;Parent=transcript1
1 Ensembl exon 1500 3000 . + . ID=exon2;Name=GENE1-001_2;Parent=transcript1
1 Ensembl exon 3500 4000 . + . ID=exon3;Name=GENE1-001_2;Parent=transcript1
1 Ensembl CDS 1300 3800 . + . ID=cds1;Name=CDS0001;Parent=transcript1
GTF format expectations
The following GTF entity types will be extracted:
cds (or CDS)
stop_codon
exon
gene
transcript
Entities are linked by an attribute named for the parent entity type e.g. exon is linked to transcript by transcript_id, transcript is linked to
gene by gene_id.
Transcript biotypes are defined in attributes named "biotype", "transcript_biotype" or "transcript_type". If none of these exist, VEP will
attempt to interpret the source field (2nd column) of the GTF as the biotype.
Here is an example:
1 Ensembl gene 1000 5000 . + . gene_id "gene1"; gene_name "GENE1";
1 Ensembl transcript 1100 4900 . + . gene_id "gene1"; transcript_id "transcript1"; gene_name
"GENE1"; transcript_name "GENE1-001"; transcript_biotype "protein_coding";
1 Ensembl exon 1200 1300 . + . gene_id "gene1"; transcript_id "transcript1"; exon_number
"exon1"; exon_id "GENE1-001_1";
1 Ensembl exon 1500 3000 . + . gene_id "gene1"; transcript_id "transcript1"; exon_number
"exon2"; exon_id "GENE1-001_2";
1 Ensembl exon 3500 4000 . + . gene_id "gene1"; transcript_id "transcript1"; exon_number
"exon3"; exon_id "GENE1-001_2";
1 Ensembl CDS 1300 3800 . + . gene_id "gene1"; transcript_id "transcript1"; exon_number
"exon2"; ccds_id "CDS0001";
Chromosome synonyms
If the chromosome names used in your GFF/GTF differ from those used in the FASTA or your input VCF, you may see warnings like this
when running VEP:
WARNING: Chromosome 21 not found in annotation sources or synonyms on line 160

To circumvent this you may provide VEP with a synonyms file. A synonym file is included in VEP's cache files, so if you have one of
these for your species you can use it as follows:
./vep -i input.vcf -cache -gff data.gff.gz -fasta genome.fa.gz -synonyms
~/.vep/homo_sapiens/112_GRCh38/chr_synonyms.txt
FASTA files
By pointing VEP to a FASTA file (or directory containing several files), it is possible to retrieve reference sequence locally when using --
cache or --offline. This enables VEP to:
Retrieve HGVS notations (--hgvs)
Check the reference sequence given in input data (--check_ref)
Construct transcript models from a GFF or GTF file without accessing a database (specially useful for performance reasons or if
using data from species/assembly not part of Ensembl species list)
FASTA files from Ensembl can be set up using the installer; files set up using the installer are automatically detected by VEP when using
--cache or --offline; you should not need to use --fasta to manually specify them.
To enable this, VEP uses one of two modules:
The Bio::DB::HTS Perl XS module with HTSlib. This module uses compiled C code and can access compressed (bgzipped) or
uncompressed FASTA files. It is set up by the VEP installer.
The Bio::DB::Fasta module. This may be used on systems where installation of the Bio::DB::HTS module has not been possible. It
can access only uncompressed FASTA files. It is also set up by the VEP installer and comes as part of the BioPerl package.
The first time you run VEP with a specific FASTA file, an index will be built. This can take a few minutes, depending on the size of the
FASTA file and the speed of your system. On subsequent runs the index does not need to be rebuilt (if the FASTA file has been modified,
VEP will force a rebuild of the index).
FASTA FTP directories
Suitable reference FASTA files are available to download from the Ensembl FTP server. See the Downloads page for details.
You should preferably use the installer as described above to fetch these files; manual instructions are provided for reference. In most
cases it is best to download the single large "primary_assembly" file for your species. You should use the unmasked (without _rm or _sm
in the name) sequences.
Note that VEP requires that the file be either unzipped (Bio::DB::Fasta) or unzipped and then recompressed with bgzip
(Bio::DB::HTS::Faidx) to run; when unzipped these files can be very large (25GB for human). An example set of commands for
setting up the data for human follows:
curl -O
https://ftp.ensembl.org/pub/release-112/fasta/homo_sapiens/dna/Homo_sapiens.GRCh38.dna.primary_ass
embly.fa.gz
gzip -d Homo_sapiens.GRCh38.dna.primary_assembly.fa.gz
bgzip Homo_sapiens.GRCh38.dna.primary_assembly.fa
./vep -i input.vcf --offline --hgvs --fasta Homo_sapiens.GRCh38.dna.primary_assembly.fa.gz
Databases
VEP can use remote or local database servers to retrieve annotations.
Using --cache (without --offline) uses the local cache on disk to fetch most annotations, but allows database connections for some
features (see cache limitations)
Using --database tells VEP to retrieve all annotations from the database. Please only use this for small input files or when using

a local database server!
Public database servers
By default, VEP is configured to connect to the public Ensembl MySQL instance at ensembldb.ensembl.org. If you are in the USA (or
geographically closer to the east coast of the USA than to the Ensembl data centre in Cambridge, UK), a mirror server is available at
useastdb.ensembl.org. To use the mirror, use the flag --host useastdb.ensembl.org
Data for Ensembl Genomes species (e.g. plants, fungi, microbes) is available through a different public MySQL server. The appropriate
connection parameters can be automatically loaded by using the flag --genomes
If you have a very small data set (100s of variants), using the public database servers should provide adequate performance. If you have
larger data sets, or wish to use VEP in a batch manner, consider one of the alternatives below.
Using a local database
It is possible to set up a local MySQL mirror with the databases for your species of interest installed. For instructions on installing a local
mirror, see here. You will need a MySQL server that you can connect to from the machine where you will run VEP (this can be the same
machine). For most of the functionality of VEP, you will only need the Core database (e.g. homo_sapiens_core_112_38) installed. In
order to find co-located variants or to use SIFT or PolyPhen, it is also necessary to install the relevant variation database (e.g.
homo_sapiens_variation_112_38).
Note that unless you have custom data to insert in the database, in most cases it will be much more efficient to use a pre-built cache in
place of a local database.
To connect to your mirror, you can either set the connection parameters using --host, --port, --user and --password, or use a registry file.
Registry files contain all the connection parameters for your database, as well as any species aliases you wish to set up:
use Bio::EnsEMBL::DBSQL::DBAdaptor;
use Bio::EnsEMBL::Variation::DBSQL::DBAdaptor;
use Bio::EnsEMBL::Registry;
Bio::EnsEMBL::DBSQL::DBAdaptor->new(
'-species' => "Homo_sapiens",
'-group' => "core",
'-port' => 5306,
'-host' => 'ensembldb.ensembl.org',
'-user' => 'anonymous',
'-pass' => '',
'-dbname' => 'homo_sapiens_core_112_38'
);
Bio::EnsEMBL::Variation::DBSQL::DBAdaptor->new(
'-species' => "Homo_sapiens",
'-group' => "variation",
'-port' => 5306,
'-host' => 'ensembldb.ensembl.org',
'-user' => 'anonymous',
'-pass' => '',
'-dbname' => 'homo_sapiens_variation_112_38'
);
Bio::EnsEMBL::Registry->add_alias("Homo_sapiens","human");
For more information on the registry and registry files, see here.
Cache - technical information
ADVANCED The cache consists of compressed files containing listrefs of serialised objects. These objects are initially created from the
database as if using the Ensembl API normally. In order to reduce the size of the cache and allow the serialisation to occur, some
changes are made to the objects before they are dumped to disk. This means that they will not behave in exactly the same way as an
object retrieved from the database when writing, for example, a plugin that uses the cache.
The following hash keys are deleted from each transcript object:
analysis
created_date
dbentries : this contains the external references retrieved when calling $transcript->get_all_DBEntries(); hence this call on a cached
object will return no entries
description
display_xref
edits_enabled
external_db
external_display_name
external_name
external_status
is_current
modified_date
status
transcript_mapper : used to convert between genomic, cdna, cds and protein coordinates. A copy of this is cached separately by
VEP as
$transcript->{_variation_effect_feature_cache}->{mapper}
As mentioned above, a special hash key "_variation_effect_feature_cache" is created on the transcript object and used to cache things
used by VEP in predicting consequences, things which might otherwise have to be fetched from the database. Some of these are stored
in place of equivalent keys that are deleted as described above. The following keys and data are stored:
introns : listref of intron objects for the transcript. The adaptor, analysis, dbID, next, prev and seqname keys are stripped from each
intron object
translateable_seq : as returned by
$transcript->translateable_seq
mapper : transcript mapper as described above
peptide : the translated sequence as a string, as returned by
$transcript->translate->seq
protein_features : protein domains for the transcript's translation as returned by
$transcript->translation->get_all_ProteinFeatures
Each protein feature is stripped of all keys but: start, end, analysis, hseqname
codon_table : the codon table ID used to translate the transcript, as returned by
$transcript->slice->get_all_Attributes('codon_table')->[0]
protein_function_predictions : a hashref containing the keys "sift" and "polyphen"; each one contains a protein function prediction
matrix as returned by e.g.
$protein_function_prediction_matrix_adaptor->fetch_by_analysis_translation_md5('sift',
md5_hex($transcript-{_variation_effect_feature_cache}->{peptide}))
Similarly, some further data is cached directly on the transcript object under the following keys:
_gene : gene object. This object has all keys but the following deleted: start, end, strand, stable_id

_gene_symbol : the gene symbol
_ccds : the CCDS identifier for the transcript
_refseq : the "NM" RefSeq mRNA identifier for the transcript
_protein : the Ensembl stable identifier of the translation
_source_cache : the source of the transcript object. Only defined in the merged cache (values: Ensembl, RefSeq) or when using a
GFF/GTF file (value: short name or filename)
!

Variant Effect Predictor Filtering results
The VEP package includes a tool, filter_vep, to filter results files on a variety of attributes.
It operates on standard, tab-delimited or VCF formatted output (NB only VCF output produced by VEP or in the same format can be
used).
Running filter_vep
Run as follows:
./vep -i in.vcf -o out.txt -cache -everything
./filter_vep -i out.txt -o out_filtered.txt -filter "[filter_text]"
filter_vep can also read from STDIN and write to STDOUT, and so may be used in a UNIX pipe:
./vep -i in.vcf -o stdout -cache -check_existing | ./filter_vep -filter "not Existing_variation" -
o out.txt
The above command removes known variants from the output
Options
Flag Alternate Description
--
help
-h
Print usage message and exit
--
inpu
t_fi
le
[fil
e]
-i
Specify the input file (i.e. the VEP results file). If no input file is specified, filter_vep
will attempt to read from STDIN. Input may be gzipped - to read a gzipped file use
--gz
--
form
at
[for
mat]
! Specify input file format:
tab (i.e. the VEP results file)
vcf
--
outp
ut_f
ile
[fil
e]
-o
Specify the output file to write to. If no output file is specified, the filter_vep will
write to STDOUT
--
forc
e_ov
erwr
ite
! Force an output file of the same name to be overwritten
--
filt
er
[fil
ters
]
-f
Add filter (see below). Multiple --filter flags may be used, and are treated as
logical ANDs, i.e. all filters must pass for a line to be printed

--
soft
_fil
ter
Variants not passing given filters will be flagged in the FILTER column of the VCF
file, and will not be removed from output.
--
list
-l
List allowed fields from the input file
--
coun
t
-c
Print only a count of matched lines
--
only
_mat
ched
! In VCF files, the CSQ field that contains the consequence data will often contain
more than one "block" of consequence data, where each block corresponds to a
variant/feature overlap. Using --only_matched will remove blocks that do not
pass the filters. By default, filter_vep prints out the entire VCF line if any of the
blocks pass the filters.
--
vcf_
info
_fie
ld
[key
]
! With VCF input files, by default filter_vep expects to find VEP annotations encoded
in the CSQ INFO key; VEP itself can be configured to write to a different key (with
the equivalent --vcf_info_field flag).
Use this flag to change the INFO key VEP expects to decode:
e.g. use the command "--vcf_info_field ANN" if the VEP annotations are
stored in the INFO key "ANN".
--
onto
logy
-y
Use Sequence Ontology to match consequence terms. Use with operator "is" to
match against all child terms of your value. e.g. "Consequence is
coding_sequence_variant" will match missense_variant, synonymous_variant etc.
Requires database connection; defaults to connecting to ensembldb.ensembl.org.
Use --host, --port, --user, --password, --version as per vep to change
connection parameters.
Writing filters
Filter strings consist of three components that must be separated by whitespace:
1.
Field : A field name from the VEP results file. This can be any field in the "main" columns of the output, or any in the "Extra" final
column. For VCF files, this is any field defined in the "##INFO=<ID=CSQ" header. You can list available fields using --list. Field
names are not case sensitive, and you may use the first few characters of a field name if they resolve uniquely to one field name.
2.
Operator : The operator defines the comparison carried out.
3.
Value : The value to which the content of the field is compared. May be prefixed with "#" to represent the value of another field.
Examples:
# match entries where Feature (Transcript) is "ENST00000307301"
--filter "Feature is ENST00000307301"
# match entries where Protein_position is less than 10
--filter "Protein_position < 10"
# match entries where Consequence contains "stream" (this will match upstream and downstream)
--filter "Consequence matches stream"
For certain fields you may only be interested in whether a value exists for that field; in this case the operator and value can be left out:
# match entries where the gene symbol is defined
--filter "SYMBOL"
The value component may be another field; to represent this, prefix the name of the field to be used as a value with "#":

# match entries where AFR_AF is greater than EUR_AF
--filter "AFR_AF > #EUR_AF"
Filter strings can be linked together by the logical operators "or" and "and", and inverted by prefixing with "not":
# filter for missense variants in CCDS transcripts where the variant falls in a protein domain
--filter "Consequence is missense_variant and CCDS and DOMAINS"
# find variants where the allele frequency is greater than 10% in either AFR or EUR populations
--filter "AFR_AF > 0.1 or EUR_AF > 0.1"
# filter out known variants
--filter "not Existing_variation"
Filter logic may be constrained using parentheses, to any arbitrary level:
# find variants with AF > 0.1 in AFR or EUR but not EAS or SAS
--filter "(AFR_AF > 0.1 or EUR_AF > 0.1) and (EAS_AF < 0.1 and SAS_AF < 0.1)"
For fields that contain string and number components, filter_vep will try and match the relevant part based on the operator in use. For
example, using --sift b in VEP gives strings that look like "tolerated(0.46)". This will give a match to either of the following filters:
# match string part
--filter "SIFT is tolerated"
# match number part
--filter "SIFT < 0.5"
Note that for numeric fields, such as the *AF allele frequency fields, filter_vep does not consider the absence of a value for that field as
equivalent to a 0 value. For example, if you wish to find rare variants by finding those where the allele frequency is less than 1% or
absent, you should use the following:
--filter "AF < 0.01 or not AF"
For the Consequence field it is possible to use the Sequence Ontology to match terms ontologically; for example, to match all coding
consequences (e.g. missense_variant, synonymous_variant):
--ontology --filter "Consequence is coding_sequence_variant"
Operators
is (synonyms: = , eq) : Match exactly
# get only transcript consequences
--filter "Feature_type is Transcript"
!= (synonym: ne) : Does not match exactly
# filter out tolerated SIFT predictions
--filter "SIFT != tolerated"
match (synonyms: matches , re , regex) : Match string using regular expression. You may include any regular expression notation,
e.g. "\d" for any numerical character
# match stop_gained, stop_lost and stop_retained
--filter "Consequence match stop"

< (synonym: lt) : Less than. Note an absent value is not considered to be equivalent to 0.
# find SIFT scores less than 0.1
--filter "SIFT < 0.1"
> (synonym: gt) : Greater than
# find variants not in the first exon
--filter "Exon > 1"
<= (synonym: lte) : Less than or equal to. Note an absent value is not considered to be equivalent to 0.
>= (synonym: gte) : Greater than or equal to
exists (synonyms: ex , defined) : Field is defined - equivalent to using no operator and value
in : Find in list or file. Value may be either a comma-separated list or a file containing values on separate lines. Each list item is
compared using the "is" operator.
# find variants in a list of gene names
--filter "SYMBOL in BRCA1,BRCA2"
# filter using a file of MotifFeatures
--filter "Feature in /data/files/motifs_list.txt"
!

Variant Effect Predictor Custom annotations
VEP can integrate custom annotation from standard format files into your results by using the --custom flag.
These files may be hosted locally or remotely, with no limit to the number or size of the files. The files must be indexed using the tabix
utility (BED, GFF, GTF, VCF); bigWig files contain their own indices.
Annotations typically appear as key=value pairs in the Extra column of the VEP output; they will also appear in the INFO column if using
VCF format output. The value for a particular annotation is defined as the identifier for each feature; if not available, an identifier derived
from the coordinates of the annotation is used. Annotations will appear in each line of output for the variant where multiple lines exist.
Data formats
VEP supports the following annotation formats:
Format Type Description Notes
GFF
GTF
Gene/transcript
annotations
Formats to describe genes and other
genomic features — format specifications:
GFF3 and GTF
Requires a FASTA file in offline mode or if the desired
species or assembly is not part of the Ensembl species
list.
VCF Variant data A format used to describe genomic variants VEP uses the 3rd column as the identifier. INFO and
FILTER fields from records may be added to the VEP
output.
BED Basic/uninterpreted
data
A simple tab-delimited format containing 3-
12 columns of data. The first 3 columns
contain the coordinates of the feature.
VEP uses the 4th column (if available) as the feature
identifier.
bigWig Basic/uninterpreted
data
A format for storage of dense continuous
data.
VEP uses the value for the given position as the identifier.
BigWig files contain their own indices, and do not need to
be indexed by tabix. Requires Bio::DB::BigFile.
Any other files can be easily converted to be compatible with VEP; the easiest format to produce is a BED-like file containing coordinates
and an (optional) identifier:
chr1 10000 11000 Feature1
chr3 25000 26000 Feature2
chrX 99000 99001 Feature3
Chromosomes can be denoted by either e.g. "chr7" or "7", "chrX" or "X".
Preparing files
Custom annotation files must be prepared in a particular way in order to work with tabix and therefore with VEP. Files must be stripped of
comment lines, sorted in chromosome and position order, compressed using bgzip and finally indexed using tabix. Here are some
examples of that process for:
GFF file
grep -v "#" myData.gff | sort -k1,1 -k4,4n -k5,5n -t$'\t' | bgzip -c > myData.gff.gz
tabix -p gff myData.gff.gz
BED file
grep -v "#" myData.bed | sort -k1,1 -k2,2n -k3,3n -t$'\t' | bgzip -c > myData.bed.gz
tabix -p bed myData.bed.gz
The tabix utility has several preset filetypes that it can process, and it can also process any arbitrary filetype containing at least a
chromosome and position column. See the documentation for details.
If you are going to use the file remotely (i.e. over HTTP or FTP protocol), you should ensure the file is world-readable on your server.

Options
Since VEP 110, you can configure each custom file using a comma-separated list of key-value pairs:
./vep [...] --custom
file=Filename,short_name=Short_name,format=File_type,type=Annotation_type,fields=VCF_fields
The order of the options is irrelevant and most options have sensible defaults as described below:
Option Accepted values Description
file
String with valid path to
file
(Required) Filename: The path to the file. For Tabix indexed files, VEP will check if both the file and
the corresponding index (.tbi) exist. For remote files, VEP will check that the tabix index is
accessible on startup.
forma
t
bed, gff, gtf, vcf or
bigwig
(Required) File format of file.
short
_name
Annotation filename
(default) or any string
without commas
Short name: A name for the annotation that will appear as the key in the key=value pairs in the
results. If not defined, this will default to the annotation filename.
field
s
VCF fields: Percentage (%) separated list of INFO fields to print (such as AC) present in the custom
input VCF or specify FILTER for the FILTER field, to add these as custom annotations:
If using exact annotation type, allele-specific annotation will be retrieved.
The INFO field name will be prefixed with the short name, e.g. using short name test, the
INFO field foo will appear as test_FOO in the VEP output. Similarly FILTER field will appear
as test_FILTER.
In VCF files the custom annotations are added to the CSQ INFO field.
Alleles in the input and VCF entry are trimmed in both directions in an attempt to match
complex or poorly formatted entries.
type
overlap (default),
within, surrounding
or exact
Annotation type:
overlap: reports any annotation that overlaps the variant by even 1 base pair.
within (*): only reports annotations within the variant.
surrounding (*): only reports annotations that completely surround the variant.
exact: only reports annotations whose coordinates match exactly those of the variant. This is
suitable for position-specific information such as conservation scores, allele frequencies or
phenotype information.
overl
ap_cu
toff
From 0 (default) to 100 Minimum percentage overlap (*) between annotation and variant. See also reciprocal.
recip
rocal
0 (default) or 1 Mode of calculating the overlap percentage (*):
0: percentage of annotation covered by variant
1: percentage of variant covered by annotation
dista
nce
0 or a positive integer
(disabled by default)
Distance (in base pairs) to the ends of the overlapping feature (*).
coord
0 (default) or 1 Force report coordinates:
Using positional options in --custom with VEP 109 and earlier (compatible with VEP 112)
Using key-value pairs in --custom with VEP 112
s
0: outputs the identifier field (or value in the case of bigWig) if available; otherwise, outputs
coordinates instead.
1: always outputs the coordinates of an overlapping custom feature.
same_
type
0 (default) or 1 Only match identical variant classes (*). For instance, only match deletions with deletions. This is
only available for VCF annotations.
num_r
ecord
s
50 (default), all, 0 or
any positive integer
Number of matching records to display. Any remaining records are represented with ellipsis
(...). Use num_records = all to display all matching records and num_records = 0 to only
display ... if there are matching records.
summa
ry_st
ats
none (default), min,
mean, max, count or
sum
Summary statistics to display. A percentage-separated list may be used to calculate multiple
summary statistics, such as min%mean%max%count%sum.
When format = vcf, the features marked with (*) only work on structural variants.
Examples:
# BigWig file
./vep [...] --custom file=frequencies.bw,short_name=Frequency,format=bigwig,type=exact,coords=0
# BED file
./vep [...] --custom
file=http://www.myserver.com/data/myPhenotypes.bed.gz,short_name=Phenotype,format=bed,type=exact,c
oords=1
# VCF file
./vep [...] --custom
file=https://ftp.ensembl.org/pub/data_files/homo_sapiens/GRCh37/variation_genotype/TOPMED_GRCh37.v
cf.gz,format=vcf,type=exact,coords=0,fields=TOPMED
./vep [...] --custom
file=gnomad_v2.1_sv.sites.vcf.gz,short_name=gnomad,fields=PC%EVIDENCE%SVTYPE,format=vcf,type=withi
n,reciprocal=1,overlap_cutoff=80
# For multiple custom files, use:
./vep [...] --custom
file=clinvar.vcf.gz,short_name=ClinVar,format=vcf,type=exact,coords=0,fields=CLNSIG%CLNREVSTAT%CLN
DN \
--custom
file=TOPMED_GRCh38_20180418.vcf.gz,short_name=topmed_20180418,format=vcf,type=exact,coords=0,field
s=TOPMED \
--custom
file=UK10K_COHORT.20160215.sites.GRCh38.vcf.gz,short_name=uk10k,format=vcf,type=exact,coords=0,fie
lds=AF_ALSPAC
Example - ClinVar
We include the most recent public variant and phenotype data available in each Ensembl release, but some projects release data more
frequently than we do.
If you want to have the very latest annotations, you can use the data files from your prefered projects (in any format listed in Data
formats) and use them as a VEP custom annotation.
For instance, you can annotate you variants with VEP, using the the latest ClinVar data as custom annotation.
ClinVar provides VCF files on their FTP site: GRCh37 and GRCh38 .
See below an example about how to use ClinVar VCF files as a VEP custom annotation:
1.
Download the VCF files (you need the compressed VCF file and the index file), e.g.:
# Compressed VCF file
curl -O https://ftp.ncbi.nlm.nih.gov/pub/clinvar/vcf_GRCh38/clinvar.vcf.gz
# Index file
curl -O https://ftp.ncbi.nlm.nih.gov/pub/clinvar/vcf_GRCh38/clinvar.vcf.gz.tbi

2.
Example of command you can use:
./vep [...] --custom
file=clinvar.vcf.gz,short_name=ClinVar,format=vcf,type=exact,coords=0,fields=CLNSIG%CLNREVSTAT%
CLNDN
## Where the selected ClinVar INFO fields (from the ClinVar VCF file) are:
# - CLNSIG: Clinical significance for this single variant
# - CLNREVSTAT: ClinVar review status for the Variation ID
# - CLNDN: ClinVar's preferred disease name for the concept specified by disease
identifiers in CLNDISDB
# Of course you can select the INFO fields you want in the ClinVar VCF file
# Quick example on GRCh38:
./vep --id "1 230710048 230710048 A/G 1" --species homo_sapiens -o /path/to/output/output.txt
--cache --offline --assembly GRCh38 --custom
file=/path/to/custom_files/clinvar.vcf.gz,short_name=ClinVar,format=vcf,type=exact,coords=0,fie
lds=CLNSIG%CLNREVSTAT%CLNDN
Using remote files
The tabix utility makes it possible to read annotation files from remote locations, for example over HTTP or FTP protocols.
In order to do this, the .tbi index file is downloaded locally (to the current working directory) when VEP is run. From this point on, only the
portions of data requested by VEP (i.e. those overlapping the variants in your input file) are downloaded.
bigWig files can also be used remotely in the same way as tabix-indexed files, although less stringent checks are carried out on VEP
startup.
!
Results in the default VEP format
Results in VCF (adding the tag --vcf in the command line)

Pathogenicity
predictions
Conservation
Pathogenicity
predictions
Conservation
Pathogenicity
predictions
Pathogenicity
predictions
Pathogenicity
predictions
Pathogenicity
Variant Effect Predictor Plugins
VEP can use plugin modules written in Perl to add functionality to the software.
Plugins are a powerful way to extend, filter and manipulate the VEP output.
They can be installed using VEP's installer script, run the following command to get a list of available plugins:
perl INSTALL.pl -a p -g list
Alternatively, VEP plugins and their dependencies are available in the Docker image. Read how to use Ensembl VEP in Docker and
Singularity.
Some plugins are also available to use via the VEP web and REST interfaces.
Existing plugins
We have written several plugins that implement experimental functionalities that we do not (yet) include in the variation API, and these
are stored in a public github repository:
https://github.com/Ensembl/VEP_plugins
Here is the list of the VEP plugins available:
Select categories:
All categories
Plugin Description Category External
libraries
Developer
AlphaMissens
e
This plugin for the Ensembl Variant Effect Predictor (VEP)
annotates missense variants with the pre-computed
AlphaMissense pathogenicity scores. AlphaMissense is a deep
learning model developed by Google DeepMind that predicts the
pathogenicity of single nucleotide missense variants. more
- Ensembl
AncestralAllel
e
A VEP plugin that retrieves ancestral allele sequences from a
FASTA file. more
- Ensembl
BayesDel This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
adds the BayesDel scores to VEP output. more
- Ensembl
Blosum62 This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
looks up the BLOSUM 62 substitution matrix score for the
reference and alternative amino acids predicted for a missense
mutation. It adds one new entry to the VEP's Extra column,
BLOSUM62 which is the associated score. more
- Ensembl
CADD
Combined
Annotation
Dependent
Depletion
A VEP plugin that retrieves CADD scores for variants from one or
more tabix-indexed CADD data files. more
- Ensembl
CAPICE A VEP plugin that retrieves CAPICE scores for variants from one
or more tabix-indexed CAPICE data files, in order to predict their
pathogenicity. more
- Ensembl
Carol A VEP plugin that calculates the Combined Annotation scoRing
toOL (CAROL) score (1) for a missense mutation based on the
pre-calculated SIFT (2) and PolyPhen-2 (3) scores from the
Ensembl API (4). more
Math::CDF
qw(pnorm
qnorm)
Ensembl

Pathogenicity
predictions
Pathogenicity
predictions
Conservation
Pathogenicity
predictions
Splicing
predictions
Variant data
Phenotype
data and
citations
Gene
tolerance to
change
Nearby
features
Visualisation
Regulatory
impact
Pathogenicity
predictions
Pathogenicity
predictions
ClinPred This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
adds pre-calculated scores from ClinPred. ClinPred is a prediction
tool to identify disease-relevant nonsynonymous variants. more
- Ensembl
Condel A VEP plugin that calculates the Consensus Deleteriousness
(Condel) score (1) for a missense mutation based on the pre-
calculated SIFT (2) and PolyPhen-2 (3) scores from the Ensembl
API (4). more
- Ensembl
Conservatio
n
This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
retrieves a conservation score from the Ensembl Compara
databases for variant positions. You can specify the method link
type and species sets as command line options, the default is to
fetch GERP scores from the EPO 35 way mammalian alignment
(please refer to the Compara documentation for more details of
available analyses). more
Net::FTP Ensembl
dbNSFP A VEP plugin that retrieves data for missense variants from a
tabix-indexed dbNSFP file. more
File::Basenam
e
qw(basename)
Ensembl
dbscSNV A VEP plugin that retrieves data for splicing variants from a tabix-
indexed dbscSNV file. more
- Ensembl
DeNovo A VEP plugin that identifies de novo variants in a VCF file. The
plugin is not compatible with JSON output format. more
List::MoreUtil
s qw(uniq)
Cwd
Ensembl
DisGeNET This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
adds Variant-Disease-PMID associations from the DisGeNET
database. It is available for GRCh38. more
List::MoreUtil
s qw(uniq)
Ensembl
DosageSensiti
vity
A VEP plugin that retrieves haploinsufficiency and triplosensitivity
probability scores for affected genes from a dosage sensitivity
catalogue published in paper -
https://www.sciencedirect.com/science/article/pii/S0092867422007
887 more
- Ensembl
Downstrea
m
This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
predicts the downstream effects of a frameshift variant on the
protein sequence of a transcript. It provides the predicted
downstream protein sequence (including any amino acids
overlapped by the variant itself), and the change in length relative
to the reference protein. more
- Ensembl
Draw A VEP plugin that draws pictures of the transcript model showing
the variant location. more
GD::Polygo
n
GD
Ensembl
Enformer This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
adds pre-calculated Enformer predictions of variant impact on
chromatin and gene expression. more
- Ensembl
EVE This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
adds information from EVE (evolutionary model of variant effect).
more
- Ensembl
FATHMM A VEP plugin that gets FATHMM scores and predictions for
missense variants. more
- Ensembl

Pathogenicity
predictions
External ID
Motif
Phenotype
data and
citations
Splicing
predictions
Phenotype
data and
citations
Frequency
data
Phenotype
data and
citations
Phenotype
data and
citations
HGVS
Functional
effect
Variant data
Look up
Gene
FATHMM_MK
L
A VEP plugin that retrieves FATHMM-MKL scores for variants from
a tabix-indexed FATHMM-MKL data file. more
- Ensembl
FlagLRG A VEP plugin that retrieves the LRG ID matching either the RefSeq
or Ensembl transcript IDs. more
Text::CSV Stephen
Kazakoff
FunMotifs This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
adds tissue-specific transcription factor motifs from FunMotifs to
VEP output. more
- Ensembl
G2P
gene2phenotype
A VEP plugin that uses G2P allelic requirements to assess
variants in genes for potential phenotype involvement. more
List::Util
qw(any)
Text::CSV
Scalar::Util
qw(looks_lik
e_number)
FileHandle
Cwd
Ensembl
GeneSplicer This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
runs GeneSplicer (https://ccb.jhu.edu/software/genesplicer/) to get
splice site predictions. more
Digest::MD5
qw(md5_hex)
Ensembl
Geno2MP A VEP plugin that adds information from Geno2MP, a web-
accessible database of rare variant genotypes linked to phenotypic
information. more
- Ensembl
gnomADc A VEP plugin that retrieves gnomAD annotation from either the
genome or exome coverage files, available here:
https://gnomad.broadinstitute.org/downloads more
File::Spec
File::Basena
me
Stephen
Kazakoff
GO
Gene Ontology
A VEP plugin that retrieves Gene Ontology (GO) terms associated
with transcripts (e.g. GRCh38) or their translations (e.g. GRCh37)
using custom GFF annotation containing GO terms. more
- Ensembl
GWAS A VEP plugin that retrieves relevant NHGRI-EBI GWAS Catalog
data given the file. more
Storable
qw(dclone)
File::Basena
me
Ensembl
HGVSIntronOf
fset
A VEP plugin for the Ensembl Variant Effect Predictor (VEP) that
returns HGVS intron start and end offsets. To be used with --hgvs
option. more
- Stephen
Kazakoff
IntAct A VEP plugin that retrieves molecular interaction data for variants
as reprted by IntAct database. more
- Ensembl
LD
Linkage
Disequilibrium
A VEP plugin that finds variants in linkage disequilibrium with any
overlapping existing variants from the Ensembl variation
databases. more
- Ensembl
LocalID The LocalID plugin allows you to use variant IDs as input without
making a database connection. more
- Ensembl

Gene
tolerance to
change
Pathogenicity
predictions
Variant data
Phenotype
data and
citations
Functional
effect
Splicing
predictions
Pathogenicity
predictions
Pathogenicity
predictions
Protein
annotation
Nearby
features
Nearby
features
LOEUF This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
adds the LOEUF scores to VEP output. LOEUF stands for the
"loss-of-function observed/expected upper bound fraction."
more
Scalar::Util
qw(looks_like_
number)
Ensembl
LoFtool
Loss-of-function
Add LoFtool scores to the VEP output. more
DBI Ensembl
LOVD
Leiden Open
Variation
Database
A VEP plugin that retrieves LOVD variation data from
http://www.lovd.nl/. more
LWP::UserAge
nt
Ensembl
Mastermind This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
uses the Mastermind Genomic Search Engine
(https://www.genomenon.com/mastermind) to report variants that
have clinical evidence cited in the medical literature. It is available
for both GRCh37 and GRCh38. more
- Ensembl
MaveDB A VEP plugin that retrieves data from MaveDB
(https://www.mavedb.org), a database that contains multiplex
assays of variant effect, including deep mutational scans and
massively parallel report assays. more
Bio::SeqUtil
s
File::Basena
me
Ensembl
MaxEntScan This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
runs MaxEntScan
(http://hollywood.mit.edu/burgelab/maxent/Xmaxentscan_scoreseq
.html) to get splice site predictions. more
Digest::MD5
qw(md5_hex)
Ensembl
MPC
missense
deleteriousness
metric
A VEP plugin that retrieves MPC scores for variants from a tabix-
indexed MPC data file. more
- Ensembl
MTR
Missense
Tolerance Ratio
A VEP plugin that retrieves Missense Tolerance Ratio (MTR)
scores for variants from a tabix-indexed flat file. more
-
Slave
Petrovski
Michael Silk
mutfunc A VEP plugin that retrieves data from mutfunc db predicting
destabilization of protein structure, interaction interface, and motif.
more
List::MoreUtil
s
qw(first_inde
x)
Compress::Z
lib
Digest::MD
5
qw(md5_hex
)
DBI
Ensembl
NearestExonJ
B
This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
finds the nearest exon junction boundary to a coding sequence
variant. More than one boundary may be reported if the
boundaries are equidistant. more
- Ensembl
NearestGen
e
This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
finds the nearest gene(s) to a non-genic variant. More than one
gene may be reported if the genes overlap the variant or if genes
are equidistant. more
- Ensembl

Protein data
Transcript
annotation
Variant data
Pathogenicity
predictions
Phenotype
data and
citations
Phenotype
data and
citations
Gene
tolerance to
change
Pathogenicity
predictions
Phenotype
data and
citations
Pathogenicity
predictions
Sequence
Sequence
neXtProt This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
retrieves data for missense and stop gain variants from neXtProt,
which is a comprehensive human-centric discovery platform that
offers integration of and navigation through protein-related data for
example, variant information, localization and interactions
(https://www.nextprot.org/). more
JSON::XS Ensembl
NMD This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
predicts if a variant allows the transcript escape nonsense-
mediated mRNA decay based on certain rules. more
- Ensembl
OpenTarget
s
A VEP plugin that integrates data from Open Targets Genetics
(https://genetics.opentargets.org), a tool that highlights variant-
centric statistical evidence to allow both prioritisation of candidate
causal variants at trait-associated loci and identification of
potential drug targets. more
Bio::SeqUtil
s
File::Basena
me
Ensembl
Paralogues A VEP plugin that fetches variants overlapping the genomic
coordinates of amino acids aligned between paralogue proteins.
This is useful to predict the pathogenicity of variants in paralogue
positions. more
Bio::SimpleA
lign
List::Util
qw(any)
File::Basena
me
Ensembl
PhenotypeOrt
hologous
A VEP plugin that retrieves phenotype information associated with
orthologous genes from model organisms. more
- Ensembl
Phenotypes A VEP plugin that retrieves overlapping phenotype information.
more
- Ensembl
pLI A VEP plugin that adds the probabililty of a gene being loss-of-
function intolerant (pLI) to the VEP output. more
List::MoreUtil
s qw/zip/
DBI
Ensembl
PON_P2 This plugin for Ensembl Variant Effect Predictor (VEP) computes
the predictions of PON-P2 for amino acid substitutions in human
proteins. more
-
Abhishek
Niroula
Mauno
Vihinen
PostGAP A VEP plugin that retrieves data for variants from a tabix-indexed
PostGAP file (1-based file). more
- Ensembl
PrimateAI The PrimateAI VEP plugin is designed to retrieve clinical impact
scores of variants, as described in
https://www.nature.com/articles/s41588-018-0167-z. Please
consider citing the paper if using this plugin. more
- Ensembl
ProteinSeqs This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
prints out the reference and mutated protein sequences of any
proteins found with non-synonymous mutations in the input file.
more
- Ensembl
ReferenceQua
lity
This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
reports on the quality of the reference genome using GRC data at
the location of your variants. More information can be found at:
- Ensembl

Pathogenicity
predictions
Transcript
annotation
Variant data
Phenotype
data and
citations
HGVS
Splicing
predictions
Splicing
predictions
Splicing
predictions
Structural
variant data
Variant data
Transcript
annotation
Nearby
features
Transcript
annotation
Pathogenicity
predictions
https://www.ncbi.nlm.nih.gov/grc/human/issues more
REVEL This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
adds the REVEL score for missense variants to VEP output.
more
- Ensembl
RiboseqORF
s
This is a VEP plugin that uses a standardized catalog of human
Ribo-seq ORFs to re-calculate consequences for variants located
in these translated regions. more
- Ensembl
SameCodon A VEP plugin that reports existing variants that fall in the same
codon. This plugin requires a database connection, can not be run
in offline mode more
- Ensembl
satMutMPR
A
A VEP plugin that retrieves data for variants from a tabix-indexed
satMutMPRA file (1-based file). The saturation mutagenesis-based
massively parallel reporter assays (satMutMPRA) measures
variant effects on gene RNA expression for 21 regulatory elements
(11 enhancers, 10 promoters). more
- Ensembl
SingleLetterA
A
This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
returns a HGVSp string with single amino acid letter codes
more
- Ensembl
SpliceAI A VEP plugin that retrieves pre-calculated annotations from
SpliceAI. SpliceAI is a deep neural network, developed by
Illumina, Inc that predicts splice junctions from an arbitrary pre-
mRNA transcript sequence. more
List::Util
qw(max)
Ensembl
SpliceRegio
n
This is a plugin for the Ensembl Variant Effect Predictor (VEP) that
provides more granular predictions of splicing effects. more
- Ensembl
SpliceVault A VEP plugin that retrieves SpliceVault data to predict exon-
skipping events and activated cryptic splice sites based on the
most common mis-splicing events around a splice site. more
- Ensembl
StructuralVari
antOverlap
A VEP plugin that retrieves information from overlapping structural
variants. more
- Ensembl
SubsetVCF A VEP plugin to retrieve overlapping records from a given VCF file.
Values for POS, ID, and ALT, are retrieved as well as values for
any requested INFO field. Additionally, the allele number of the
matching ALT is returned. more
Data::Dumpe
r
Storable
qw(dclone)
Joseph A.
Prinz
TranscriptAnn
otator
A VEP plugin that annotates variant-transcript pairs based on a
given file: more
File::Basenam
e
Ensembl
TSSDistanc
e
A VEP plugin that calculates the distance from the transcription
start site for upstream variants. more
- Ensembl
UTRAnnotato
r
A VEP plugin that annotates the effect of 5' UTR variant especially
for variant creating/disrupting upstream ORFs. Available for both
GRCh37 and GRCh38. more
List::Util
qw(min max)
Scalar::Util
qw(looks_lik
e_number)
Ensembl
VARITY This is a plugin for the Ensembl Variant Effect Predictor (VEP) that - Ensembl

predictions
adds the pre-computed VARITY scores to predict pathogenicity of
rare missense variants to VEP output. more
We hope that these will serve as useful examples for users implementing new plugins. If you have any questions about the system, or
suggestions for enhancements please let us know on the ensembl-dev mailing list.
We also encourage you to share any plugins you develop: we are happy to accept pull requests on the VEP_plugins git repository.
There are further published plugins available outside the VEP repository including:
LOFTEE a Loss-Of-Function Transcript Effect Estimator (Konrad Karczewski et al,2020)
How it works
Plugins are run once VEP has finished its analysis for each line of the output, but before anything is printed to the output file.
When each plugin is called (using the run method) it is passed two data structures to use in its analysis; the first is a data structure
containing all the data for the current line, and the second is a reference to a variation API object that represents the combination of a
variant allele and an overlapping or nearby genomic feature (such as a transcript or regulatory region).
This object provides access to all the relevant API objects that may be useful for further analysis by the plugin (such as the current
VariationFeature and Transcript).
Please refer to the Ensembl Variation API documentation for more details.
Functionality
We expect that most plugins will simply add information to the last column of the output file, the "Extra" column, and the plugin system
assumes this in various places, but plugins are also free to alter the output line as desired.
The only hard requirement for a plugin to work with VEP is that it implements a number of required methods (such as new which should
create and return an instance of this plugin, get_header_info which should return descriptions of the type of data this plugin produces to
be included in VEP output's header, and run which should actually perform the logic of the plugin).
To make development of plugins easier, we suggest that users use the Bio::EnsEMBL::Variation::Utils::BaseVepPlugin module as their
base class, which provides default implementations of all the necessary methods which can be overridden as required.
Please refer to the documentation in this module for details of all required methods and for a simple example of a plugin implementation.
Filtering using plugins
A common use for plugins will be to filter the output in some way (for example to limit output lines to missense variants) and so we
provide a simple mechanism to support this.
The run method of a plugin is assumed to return a reference to a hash containing information to be included in the output, and if a plugin
should not add any data to a particular line it should return an empty hashref. If a plugin should instead filter a line and exclude it from
the output, it should return undef from its run method, this also means that no further plugins will be run on the line.
If you are developing a filter plugin, we suggest that you use the Bio::EnsEMBL::Variation::Utils::BaseVepFilterPlugin as your base class
and then you need only override the include_line method to return true if you want to include this line, and false otherwise.
Again, please refer to the documentation in this module for more details and an example implementation of a missense filter.
Using plugins
In order to run a plugin you need to include the plugin module in Perl's library path somehow; by default VEP includes the ~/.vep/Plugins
directory in the path, so this is a convenient place to store plugins, but you are also able to include modules by any other means (e.g
using the $PERL5LIB environment variable in Unix-like systems).
You can then run a plugin using the --plugin command line option, passing the name of the plugin module as the argument.
For example, if your plugin is in a module called MyPlugin.pm, stored in ~/.vep/Plugins, you can run it with a command line like:
./vep -i input.vcf --plugin MyPlugin
You can pass arguments to the plugin's 'new' method by including them after the plugin name on the command line, separated by
commas, e.g.:
./vep -i input.vcf --plugin MyPlugin,1,FOO

If your plugin inherits from BaseVepPlugin, you can then retrieve these parameters as a list from the params method.
You can run multiple plugins by supplying multiple --plugin arguments. Plugins are run serially in the order in which they are specified on
the command line, so they can be run as a pipeline, with, for example, a later plugin filtering output based on the results from an earlier
plugin. Note though that the first plugin to filter a line 'wins', and any later plugins won't get run on a filtered line.
Intergenic variants
When a variant falls in an intergenic region, it will usually not have any consequence types called, and hence will not have any
associated VariationFeatureOverlap objects. In this special case, VEP creates a new VariationFeatureOverlap that overlaps a feature of
type "Intergenic".
To force your plugin to handle these, you must add "Intergenic" to the feature types that it will recognize; you do this by writing your own
feature_types sub-routine:
sub feature_types {
return ['Transcript', 'Intergenic'];
}
This will cause your plugin to handle any variation features that overlap transcripts or intergenic regions. To also include any regulatory
features, you should use the generic type "Feature":
sub feature_types {
return ['Feature', 'Intergenic'];
}
!

Variant Effect Predictor Examples and use cases
Example commands
Read input from STDIN, output to STDOUT
./vep --cache -o stdout
Add regulatory region consequences
./vep --cache -i variants.txt --regulatory
Input file variants.vcf.txt, input file format VCF, add gene symbol identifiers
./vep --cache -i variants.vcf.txt --format vcf --symbol
Filter out common variants based on 1000 Genomes data
./vep --cache -i variants.txt --filter_common
Force overwrite of output file variants_output.txt, check for existing co-located variants, output only coding sequence
consequences, output HGVS names
./vep --cache -i variants.txt -o variants_output.txt --force --check_existing --coding_only --
hgvs
Run for any species or assembly (even if not part of Ensembl data) by providing your own FASTA file and GFF/GTF annotation
./vep -i variants.txt -o variants_output.txt --gff data.gff.gz --fasta genome.fa.gz
Specify DB connection parameters in registry file ensembl.registry, add SIFT score and prediction, PolyPhen prediction
./vep --database -i variants.txt --registry ensembl.registry --sift b --polyphen p
Connect to Ensembl Genomes db server for Arabidopsis thaliana
./vep --database -i variants.txt --genomes --species arabidopsis_thaliana
Load config from ini file, run in quiet mode
./vep --config vep.ini -i variants.txt -q
Use cache in /home/vep/mycache/, use gzcat instead of zcat
./vep --cache --dir /home/vep/mycache/ -i variants.txt --compress gzcat
Add custom position-based phenotype annotation from remote BED file
./vep --cache -i variants.vcf --custom
file=ftp://ftp.myhost.org/data/phenotypes.bed.gz,short_name=phenotype
Use the plugin named MyPlugin, output only the variation name, feature, consequence type and MyPluginOutput fields
./vep --cache -i variants.vcf --plugin MyPlugin --fields

Uploaded_variation,Feature,Consequence,MyPluginOutput
Right align variants before consequence calculation. For more information, see here.
./vep --cache -i variants.vcf --shift_3prime 1
Report uploaded allele before minimisation. For more information, see here.
./vep --cache -i variants.vcf --uploaded_allele
gnomAD
gnomAD exome frequency data is included in VEP's cache files from release 90, replacing ExAC; use --af_gnomade to enable using
this data. VEP can also retrieve frequency data from the gnomAD genomes set or ExAC via VEP's custom annotation functionality.
For the latest gnomAD data, please visit gnomAD downloads .
1.
VEP requires Bio::DB::HTS to read data from tabix-indexed VCFs - see installation instructions
2.
Ensembl's FTP site hosts abridged VCF files for gnomAD and ExAC, additionally remapped to GRCh38 using CrossMap . It is
possible for VEP to read these files directly from their remote location, though for optimal performance the VCF and index should be
downloaded to a local file system.
GRCh38
gnomAD genomes (r2.1, remapped with CrossMap): [VCFs and tabix indexes]
gnomAD exomes (r2.1, remapped with CrossMap): [VCFs and tabix indexes]
ExAC (v0.3, remapped using CrossMap): [VCF] [tabix index]
GRCh37
gnomAD genomes (r2.1): [VCF and tabix indexes]
gnomAD exomes (r2.1): [VCF and tabix indexes]
ExAC (v0.3): [VCF] [tabix index]
3.
Run VEP with the following command (using the GRCh38 input example) to get locations and continental-level allele frequencies:
./vep -i examples/homo_sapiens_GRCh38.vcf --cache \
--custom
file=gnomad.genomes.r2.0.1.sites.GRCh38.noVEP.vcf.gz,short_name=gnomADg,format=vcf,type=exact,c
oords=0,fields=AF_AFR%AF_AMR%AF_ASJ%AF_EAS%AF_FIN%AF_NFE%AF_OTH
You will then see data under field names as described in the VEP output header:
## gnomADg : gnomad.genomes.r2.0.1.sites.GRCh38.noVEP.vcf.gz (exact)
## gnomADg_AFR_AF : AFR_AF field from gnomad.genomes.r2.0.1.sites.GRCh38.noVEP.vcf.gz
## gnomADg_AMR_AF : AMR_AF field from gnomad.genomes.r2.0.1.sites.GRCh38.noVEP.vcf.gz
...
where the gnomADg field contains the ID (or coordinates if no ID found) of the variant in the VCF file. Any of the fields in the
gnomAD file INFO field can be added by appending them to the list in your VEP command.
Conservation scores
You can use VEP's custom annotation feature to add conservation scores to your output. For example, to add GERP scores, download
the bigWig file from the list below, and run VEP with the following flag:
./vep --cache -i example.vcf --custom file=All_hg19_RS.bw,short_name=GERP,format=bigwig

Human (GRCh38)
phastCons 7-way
phastCons 20-way
phastCons 100-way
phyloP 7-way
phyloP 20-way
phyloP 100-way
Human (GRCh37)
GERP
phastCons 46-way
phastCons 100-way
phyloP 46-way
phyloP 100-way
Example conservation score files:
All files provided by the UCSC genome browser - files for other species are available from their FTP site , though be sure to use the file
corresponding to the correct assembly.
dbNSFP
dbNSFP - "a lightweight database of human nonsynonymous SNPs and their functional predictions" - provides pathogenicity
predictions from many tools (including SIFT, LRT, MutationTaster, FATHMM) across every possible missense substitution in the human
proteome.
Plugins in VEP sometimes require data processed in specific ways as arguments. Any requirements and usage instructions for each
plugin can be found in the plugin documentation.
In the case of the dbNSFP.pm plugin, the data needs to be downloaded and then processed into a format that the plugin can use. Note
that there are two distinct branches of the files provided for academic and commercial usage; please use the appropriate files for your
use case.
After downloading the file, you will need to process it so that tabix can index it correctly. This will take a while as the file is very large!
Note that you will need the tabix utility in your path to use dbNSFP.
version=4.5c
unzip dbNSFP${version}.zip
zcat dbNSFP${version}_variant.chr1.gz | head -n1 > h
# GRCh38/hg38 data
zgrep -h -v "^#chr" dbNSFP${version}_variant.chr* | sort -k1,1 -k2,2n - | cat h - | bgzip -c >
dbNSFP${version}_grch38.gz
tabix -s 1 -b 2 -e 2 dbNSFP${version}_grch38.gz
# GRCh37/hg19 data
zgrep -h -v "^#chr" dbNSFP${version}_variant.chr* | awk '$8 != "." ' | sort -k8,8 -k9,9n - | cat h
- | bgzip -c > dbNSFP${version}_grch37.gz
tabix -s 8 -b 9 -e 9 dbNSFP${version}_grch37.gz
Then simply download the dbNSFP.pm plugin and place it either in $HOME/.vep/Plugins/ or a path in your $PERL5LIB. When you
run VEP with the plugin, you will need to select some of the columns that you wish to retrieve; to list them run VEP with the plugin and
the path to the dbNSFP file and no further parameters:
./vep --cache --force --plugin dbNSFP,dbNSFP4.5c_grch38.txt.gz
2014-04-04 11:27:05 - Read existing cache info
2014-04-04 11:27:05 - Auto-detected FASTA file in cache directory
2014-04-04 11:27:05 - Checking/creating FASTA index
2014-04-04 11:27:05 - Failed to instantiate plugin dbNSFP: ERROR: No columns selected to fetch.
Available columns are:
#chr,pos(1-coor),ref,alt,aaref,aaalt,hg18_pos(1-coor),genename,Uniprot_acc,
Uniprot_id,Uniprot_aapos,Interpro_domain,cds_strand,refcodon,SLR_test_statistic,
codonpos,fold-degenerate,Ancestral_allele,Ensembl_geneid,Ensembl_transcriptid,
...
Note that some of these fields are replicates of those produced by the core VEP code (e.g. SIFT, the 1000 Genomes and ESP
frequencies) - you should use the options to enable these from the VEP code in place of the annotations from dbNSFP as the dbNSFP
file covers only missense substitutions. Other fields, such as the conservation scores, may be better served by using genome-wide files
as described above.

To select fields, just add them as a comma-separated list to your command line:
./vep --cache --force --plugin
dbNSFP,dbNSFP4.5c_grch38.txt.gz,LRT_score,FATHM_score,MutationTaster_score
One final point to note is that the dbNSFP scores are frozen on a particular Ensembl release's transcript set; check the readme file on
their download site to find out exactly which. While in the majority of cases protein sequences don't change between releases, in some
circumstances the protein sequence used by VEP in the latest release may differ from the sequence used to calculate the scores in
dbNSFP.
Structural Variants
VEP can be used to annotate structural variants (SV) with their predicted effect on other genomic features. For more information on SV
input format, see here.
Prediction process
The INFO keys 'END' or 'SVLEN' are present, the proportion of any overlapping feature covered by the variant is calculated
If the SVTYPE or ALT is 'DEL', the variant tested for feature ablation/ truncation
If the SVTYPE or ALT is 'DUP', the variant tested for feature amplification
If the SVTYPE or ALT is 'INS' or 'DUP', the variant tested for feature elongatation
SVTYPE is used in preference to ALT to derive the variant type of an SV with 'CN*' alleles
Reported overlaps
VEP calculates the length and proportion of each genomic feature overlapped by a structural variant
Use the --overlaps option to enable this when using VCF or tab format. (This is reported by default in standard VEP and JSON
format.)
The keys bp_overlap and percentage_overlap are used in JSON format and OverlapBP and OverlapPC in other formats.
Changing memory requirements
By default, VEP does not annotate variants larger than 10M. If you are using the command line tool, you can use the --max_sv_size
option to modify this.
By default, variants are analysed in batches of 5000. Using the --buffer_size option to reduce this can reduce memory requirements,
especially if your data is sparse. A smaller buffer size is essential when annotating structural variants with regulatory data.
Citations and VEP users
VEP is used by many organisations and projects:
VEP forms a part of Illumina's VariantStudio software
Gemini is a framework for exploring genome variation that uses VEP
The DECIPHER project uses VEP in its analysis pipelines
Other citations and use cases:
VAX is a suite of plugins for VEP that expands its functionality
pViz is a visualisation tool for VEP results files
McCarthy et al compares VEP to AnnoVar
Pabinger et al reviews variant analysis software, including VEP
VEP is used to provide annotation for the ExAC and gnomAD projects
!

Variant Effect Predictor Other information
Getting VEP to run faster
Set up correctly, VEP is capable of processing around 3 million variants in 30 minutes. There are a number of steps you can take to
make sure your VEP installation is running as fast as possible:
1.
Make sure you have the latest version of VEP and the Ensembl API. We regularly introduce optimisations, alongside the new
features and bug fixes of a typical new release.
2.
Download a cache file for your species. If you are using --database, you should consider using --cache or --offline instead. Any time
VEP has to access data from the database (even if you have a local copy), it will be slower than accessing data in the cache on your
local file system.
Enabling certain flags forces VEP to access the database, and you will be warned at startup that it will do this with e.g.:
2011-06-16 16:24:51 - INFO: Database will be accessed when using --check_svs
Consider carefully whether you need to use these flags in your analysis.
3.
If you use --check_existing or any flags that invoke it (e.g. --af, --af_1kg, --filter_common, --everything), tabix-convert your cache file.
Checking for known variants using a converted cache is >100% faster than using the default format.
4.
Download a FASTA file (and use the flag --fasta) if you use --hgvs or --check_ref. Again, this will prevent VEP accessing the
database unnecessarily (in this case to retrieve genomic sequence).
5.
Using forking enables VEP to run multiple parallel "threads", with each thread processing a subset of your input. Most modern
computers have more than one processor core, so running VEP with forking enabled can give huge speed increases (3-4x faster in
most cases). Even computers with a single core will see speed benefits due to overheads associated with using object-oriented
code in Perl.
To use forking, you must choose a number of forks to use with the --fork flag. We recommend using 4 forks:
./vep -i my_input.vcf --fork 4 --offline
but depending on various factors specific to your setup you may see faster performance with fewer or more forks.
When writing plugins be aware that while the VEP code attempts to preserve the state of any plugin-specific cached data between
separate forks, there may be situations where data is lost. If you find this is the case, you should disable forking in the new() method
of your plugin by deleting the "fork" key from the $config hash.
6.
Make sure your cache and FASTA files are stored on the fastest file system or disk you have available. If you have a lot of memory
in your machine, you can even pre-copy the files to memory using tmpfs .
7.
Consider if you need to generate HGVS notations (--hgvs); this is a complex annotation step that can add ~50-80% to your runtime.
Note also that --hgvs is switched on by --everything.
8.
Install the Set::IntervalTree Perl package. This package speeds up VEP's internals by changing how overlaps between variants
and transcript components are calculated.
9.
Install the Ensembl::XS package. This contains compiled versions of certain key subroutines used in VEP that will run faster than
the default native Perl equivalents. Using this should improve runtime by 5-10%.
10.
Add the --no_stats flag. Calculating summary statistics increases VEP runtime, so can be switched off if not required
11.
VEP is optimised to run on input files that are sorted in chromosomal order. Unsorted files will still work, albeit more slowly.
12.
For very large files (for example those from whole-genome sequencing), VEP process can be easily parallelised by dividing your file
into chunks (e.g. by chromosome). VEP will also work with tabix-indexed, bgzipped VCF files, and so the tabix utility could be used
to divide the input file:
tabix -h variants.vcf.gz 12:1000000-20000000 | ./vep --cache --vcf

Species with multiple assemblies
Ensembl currently supports the two latest human assembly versions. We provide a VEP cache using the latest software version (112) for
both GRCh37 and GRCh38.
The VEP installer will install and set up the correct cache and FASTA file for your assembly of interest. If using the --AUTO functionality
to install without prompts, remember to add the assembly version required using e.g. "--ASSEMBLY GRCh37". It is also possible to have
concurrent installations of caches from both assemblies; just use the --assembly to select the correct one when you run VEP.
Once you have installed the relevant cache and FASTA file, you are then able to use VEP as normal. If you are using GRCh37 and
require database access in addition to the cache (for example, to look up variant identifiers using --format id, see cache limitations), you
will be warned you that you must change the database port in order to connect to the correct database:
ERROR: Cache assembly version (GRCh37) and database or selected assembly version (GRCh38) do not
match
If using human GRCh37 add "--port 3337" to use the GRCh37 database, or --offline to avoid database
connection entirely
If you have data you wish to map to a new assembly, you can use the Ensembl assembly converter tool - if you've downloaded VEP,
then you have it already! The tool is found in the ensembl-tools/scripts/assembly_converter folder. There is also an online version of the
tool available. Both UCSC (liftOver ) and NCBI (Remap ) also provide tools for converting data between assemblies.
Summarising annotation
By default VEP is configured to provide annotation on every genomic feature that each input variant overlaps. This means that if a
variant overlaps a gene with multiple alternate splicing variants (transcripts), then a block of annotation for each of these transcripts is
reported in the output. In the default VEP output format each of these blocks is written on a single line of output; in VCF output format the
blocks are separated by commas in the INFO field.
A number of options are provided to reduce the amount of output produced if this depth of annotation is not required.
Example
Input data (VCF - input.vcf)
##fileformat=VCFv4.2
#CHROM POS ID REF ALT
1 230710048 rs699 A G
1 230710514 var_2 A G,T
Example of VEP command and output (no "pick" option):
./vep --cache -i input.vcf -o output.txt
#Uploaded_variation Location Allele Gene Feature Feature_type Consequence cDNA_position
CDS_position Protein_position Amino_acids Codons Existing_variation Extra
rs699 1:230710048 G ENSG00000135744 ENST00000366667 Transcript missense_variant 1018
803 268 M/T aTg/aCg - IMPACT=MODERATE;STRAND=-1
rs699 1:230710048 G ENSG00000244137 ENST00000412344 Transcript downstream_gene_variant -
- - - - - IMPACT=MODIFIER;DISTANCE=650;STRAND=-1
var_2 1:230710514 G ENSG00000135744 ENST00000366667 Transcript synonymous_variant 552
337 113 L Ttg/Ctg - IMPACT=LOW;STRAND=-1
var_2 1:230710514 T ENSG00000135744 ENST00000366667 Transcript missense_variant 552
337 113 L/M Ttg/Atg - IMPACT=MODERATE;STRAND=-1
var_2 1:230710514 G ENSG00000244137 ENST00000412344 Transcript downstream_gene_variant -
- - - - - IMPACT=MODIFIER;DISTANCE=184;STRAND=-1
var_2 1:230710514 T ENSG00000244137 ENST00000412344 Transcript downstream_gene_variant -
- - - - - IMPACT=MODIFIER;DISTANCE=184;STRAND=-1
Options

--pick
VEP chooses one block of annotation per variant, using an ordered set of criteria. This order may be customised using --pick_order.
1.
MANE Select transcript status
2.
MANE Plus Clinical transcript status
3.
canonical status of transcript
4.
APPRIS isoform annotation
5.
transcript support level
6.
biotype of transcript ("protein_coding" preferred)
7.
CCDS status of transcript
8.
consequence rank according to this table
9.
translated, transcript or feature length (longer preferred)
Show example of VEP command and output, with the "--pick" option.
--pick_allele
As above, but chooses one consequence block per variant allele. This can be useful for VCF input files with more than one ALT
allele.
Show example of VEP command and output, with the "--pick_allele" option.
--per_gene
As --pick, but chooses one annotation block per gene that the input variant overlaps.
Show example of VEP command and output, with the "--per_gene" option.
--pick_allele_gene
As above, but chooses one consequence block per variant allele and gene combination.
Show example of VEP command and output, with the "--pick_allele_gene" option.
--flag_pick
Instead of choosing one block and removing the others, this option adds a flag "PICK=1" to picked annotation block, allowing you to
easily filter on this later using VEP's filtering tool.
--flag_pick_allele
As above, but flags one block per allele.
--flag_pick_allele_gene
As above, but flags one block per allele and gene combination.
--most_severe
This flag reports only the consequence type of the block with the highest rank, according to this table.
Show example of VEP command and output, with the "--most_severe" option.
--summary
This flag reports only a comma-separated list of the consequence types predicted for this variant.
Show example of VEP command and output, with the "--summary" option.

HGVS notations
Output
HGVS notations can be produced by VEP using the --hgvs flag. Coding (c.) and protein (p.) notations given against Ensembl identifiers
use versioned identifiers that guarantee the identifier refers always to the same sequence.
Genomic HGVS notations may be reported using --hgvsg. Note that the named reference for HGVSg notations will be the chromosome
name from the input (as opposed to the officially recommended chromosome accession).
HGVS notations for insertions or deletions are by default shifted 3-prime relative to the reported transcript or protein sequence in
accordance with HGVS specifications. This may lead to discrepancies between the coordinates reported in the HGVS nomenclature and
the coordinate columns reported by VEP. You may instruct VEP not to shift using --shift_hgvs 0.
Reference sequence used as part of VEP's HGVSc calculations is taken from a given FASTA file, rather than the variant reference.
HGVSp is calculated using the given variant reference.
Input
VEP supports using HGVS notations as input. This feature is currently under development and not all HGVS notation types are
supported. Notations relative to genomic (g.) or coding (c.) sequences are fully supported; protein (p.) notations are supported in limited
fashion due to the complexity involved in determining the multiple possible underlying genomic sequence changes that could produce a
single protein change. A warning will be given if a particular notation cannot be parsed.
By default VEP uses Ensembl transcripts as the reference for determining consequences, and hence also for HGVS notations. However,
it is possible to parse HGVS notations that use RefSeq transcripts as the reference sequence by using the --refseq flag. Such notations
must include the version number of the transcript e.g.
NM_080794.3:c.1001C>T
where ".3" denotes that this is version 3 of the transcript NM_080794. See below for more details on how VEP can use RefSeq
transcripts.
RefSeq transcripts
If you prefer to exclude predicted RefSeq transcripts (those with identifiers beginning with "XM_" or "XR_") use --exclude_predicted.
Identifiers and other data
VEP's RefSeq cache lacks many classes of data present in the Ensembl transcript cache.
Included in the RefSeq cache
Gene symbol
SIFT and PolyPhen predictions
Not included in the RefSeq cache
APPRIS annotation
TSL annotation
UniProt identifiers
CCDS identifiers
Protein domains
Gene-phenotype association data
Differences to the reference genome
RefSeq transcript sequences may differ from the genome sequence to which they are aligned. Ensembl's API (and hence VEP)
constructs transcript models using the genomic reference sequence. These differences are accounted for using BAM-edited transcript
models. in human cache files from release 90 onwards. Prior to release 90 and in non-human species differences between the RefSeq

sequence and the genomic sequence are not accounted for, so some annotations produced by VEP on these transcripts may be
inaccurate. Most differences occur in non-coding regions, typically in UTRs at either end of transcripts or in the addition of a poly-A tail,
causing minimal impact on annotation.
For human VEP cache files, each RefSeq transcript is annotated with the REFSEQ_MATCH flag indicating whether and how the RefSeq
model differs from the underlying genome.
Correcting transcript models with BAM files
NCBI have released BAM files that contain alignments of RefSeq transcripts to the genome. From release 90 onwards, these alignments
have been incorporated and used to correct the transcript models in the human RefSeq and merged cache files.
VEP's cache building process uses the sequence and alignment in the BAM to correct the RefSeq model. If the corrected model does
not match the original RefSeq sequence in the BAM, the corrected model is discarded. The success or failure of the BAM edit is
recorded in the BAM_EDIT field of the VEP output. Failed edits are extremely rare (< 0.01% of transcripts), but any VEP annotations
produced on transcripts with a failed edit status should be interpreted with extreme caution.
Using BAM-edited transcripts causes VEP to change how alleles are interpreted from input variants. Input variants are typically encoded
in VCFs that are called using the reference genome. This means that the alternate (ALT) allele as given in the VCF may correspond to
the reference allele as found in the corrected RefSeq transcript model. VEP will account for this, using the corrected reference allele (by
enabling --use_transcript_ref) when calculating consequences, and the GIVEN_REF and USED_REF fields in the VEP output indicate
any change made. If the reference allele derived from the transcript matches any given alternate (ALT) allele, then no consequence data
will be produced for this allele as it will be considered non-variant. Note that this process may also clash with any interpretation from
using --check_ref, so it is recommended to avoid using this flag.
To override the behaviour of --use_transcript_ref and force VEP to use your input reference allele instead of the one derived from the
transcript, you may use --use_given_ref.
VEP can also side-load BAM files at runtime to correct transcript models on-the-fly; this allows corrections to be applied for other
species, where alignments are available, or when using RefSeq GFF files, rather than the cache.
./vep --cache --refseq -i variants.vcf --species mus_musculus --bam
GCF_000001635.26_GRCm38.p6_knownrefseq_alns.bam
BAM files are available from NCBI:
Human GRCh38.p13
Human GRCh37.p13
Existing or colocated variants
Use the --check_existing flag to identify known variants colocated with input variant. VEP's known variant cache is derived from
Ensembl's variation database and contains variants from dbSNP and other sources.
VEP by default uses a normalisation-based allele matching algorithm to identify known variants that match input variants. Since both
input and known variants may have multiple alternate (ALT) or variant alleles, each pair of reference (REF) and ALT alleles are
normalised and compared independently to arrive at potential matches. VCF permits multiple allele types to be encoded on the same
line, while dbSNP assigns separate rsID identifiers to different allele types at the same locus. This means different alleles from the same
input variant may be assigned different known variant identifiers.

Illustration of VEP's allele matching algorithm resolving one VCF line with multiple ALTs to three different variant types and coordinates
Note that allele matching occurs independently of any allele transformations carried out by --minimal; VEP will match to the same
identifiers and frequency data regardless of whether the flag is used.
For some data sources (COSMIC, HGMD), Ensembl is not licensed to redistribute allele-specific data, so VEP will report the existence of
co-located variants with unknown alleles without carrying out allele matching. To disable this behaviour and exclude these variants, use
the --exclude_null_alleles flag.
To disable allele matching completely and compare variant locations only, use --no_check_alleles.
Frequency data
In addition to identifying known variants, VEP also reports allele frequencies for input alleles from major genotyping projects (1000
genomes, gnomAD exomes and gnomAD genomes). VEP's cache currently contains only frequency data for alleles that have been
submitted to dbSNP or are imported via another source into the Ensembl variation database. This means that until gnomAD's full data
set is submitted to dbSNP and incorporated into Ensembl, the frequency for some alleles may be missing from VEP's cache data.
To access the full gnomAD data set, it is possible to use VEP's custom annotation feature to retrieve the frequency data directly from the
gnomAD VCF files; see instructions here.
Normalising Consequences
Insertions and deletions in repetitive sequences can be often described at different equivalent locations and may therefore be assigned
different consequence predictions. VEP can optionally convert variant alleles to their most 3’ representation before consequence
calculation.
In the example below, we insert a G at the start of the repeated region. Without the --shift_3prime flag, VEP will calculate consequences
at the input position and report the variant as a frameshift, and recognising that the variant lies within 2 bases of a splice site, as
splice_region_variant.

./vep --cache -id '3 46358467 . A AG'
#Uploaded_variation Location Allele Gene Feature Feature_type Consequence
cDNA_position CDS_position Protein_position Amino_acids Codons Existing_variation
Extra
3_46358468_-/G 3:46358467-46358468 G ENSG00000121807 ENST00000292301 Transcript
frameshift_variant,splice_region_variant 1425-1426 940-941 314 S/RX agc/aGgc
-
IMPACT=HIGH;STRAND=1
...
However, with --shift_3prime switched on, VEP will right align all insertions and deletions within repeated regions, shifting the inserted G
two positions to the right before consequence calculation, providing the splice_donor_variant consequence instead.
./vep --cache -id '3 46358467 . A AG' --shift_3prime 1
#Uploaded_variation Location Allele Gene Feature Feature_type Consequence
cDNA_position CDS_position Protein_position Amino_acids Codons Existing_variation
Extra
3_46358468_-/G 3:46358467-46358468 G ENSG00000121807 ENST00000292301 Transcript
splice_donor_variant - - - - - - IMPACT=HIGH;STRAND=1
...
Using --shift_genomic will also update the location field. However, --shift_genomic will also shift intergenic variants, which can lead to a
reduction in performance.
./vep --cache -id '3 46358467 . A AG' --shift_genomic 1
#Uploaded_variation Location Allele Gene Feature Feature_type Consequence
cDNA_position CDS_position Protein_position Amino_acids Codons Existing_variation
Extra
3_46358468_-/G 3:46358469-46358470 G ENSG00000121807 ENST00000292301 Transcript
splice_donor_variant - - - - - - IMPACT=HIGH;STRAND=1
...
When shifting, insertions or deletions of length 2 or more can lead to alterations in the reported alternate allele. For example, an insertion
of GAC that can be shifted 2 bases in the 3' direction will alter the alternate allele to CGA.
./vep --cache -id '3 46358464 . A AGAC' --shift_3prime 1
#Uploaded_variation Location Allele Gene Feature Feature_type Consequence
cDNA_position CDS_position Protein_position Amino_acids Codons Existing_variation
Extra
3_46358465_-/GAC 3:46358464-46358465 CGA ENSG00000121807 ENST00000292301 Transcript
inframe_insertion,splice_region_variant 1424-1425 939-940 313-314 -/R -/CGA -
IMPACT=MODERATE;STRAND=1
...
./vep --cache -id '3 46358464 . A AGAC' --shift_3prime 0

#Uploaded_variation Location Allele Gene Feature Feature_type Consequence
cDNA_position CDS_position Protein_position Amino_acids Codons Existing_variation
Extra
3_46358465_-/GAC 3:46358464-46358465 GAC ENSG00000121807 ENST00000292301 Transcript
inframe_insertion 1422-1423 937-938 313 R/RR aga/aGACga -
IMPACT=MODERATE;STRAND=1
!

Variant Effect Predictor FAQ
For any questions not covered here, please send an email to the Ensembl developer's mailing list (public) or contact the Ensembl
Helpdesk (private). Also you can report issues through our (public) Github repositories. For general vep issues you should use ensembl-
vep repository and for specific plugins you should use VEP_plugins repository.
General questions
Q: Why has my insertion/deletion variant encoded in VCF disappeared from the VEP output?
Ensembl treats unbalanced variants differently to VCF - your variant hasn't disappeared, it may have just changed slightly! You can solve
this by giving your variants a unique identifier in the third column of the VCF file. See here for a full discussion.
!
Q: Why don't I see any co-located variants when using species X?
Ensembl only has variation databases for a subset of all Ensembl species - see this document for details.
!
Q: Why do I see multiple known variants mapped to my input variant?
VEP compares your input to known variants from the Ensembl variation database. In some cases one input variant can match multiple
known variants:
Germline variants from dbSNP and somatic mutations from COSMIC may be found at the same locus
Some sources, e.g. HGMD, do not provide public access to allele-specific data, so an HGMD variant with unknown alleles may
colocate with one from dbSNP with known alleles
Multiple alternate alleles from your input may match different variants as they are described in dbSNP
See here for a full discussion.
!
Q: VEP is not assigning a frequency to my input variant - why?
VEP's cache contains frequency data only for variants and alleles imported into Ensembl's variation database. See here for a full
discussion.
!
Q: Why do I see so many lines of output for each variant in my input?
While it would be convenient to have a simple, one word answer to the question "What is the consequence of this variant?", in reality
biology is not this simple! Many genes have more than one transcript, so VEP provides a prediction for each transcript that a variant
overlaps. VEP has options to help select results according to your requirements; the --canonical and --ccds options indicate which
transcripts are canonical and belong to the CCDS set respectively, while --pick, --per_gene, --summary and --most_severe allow you to
give a more summary level assessment per variant.
Furthermore, several "compound" consequences are also possible - if, for example, a variant falls in the final few bases of an exon, it
may be considered to affect a splicing site, in addition to possibly affecting the coding sequence.
!
Q: How do I reduce VEP's memory requirement?
There are a number of ways to do this-
1.
Ensure your input file is sorted by location. This can greatly reduce memory requirements and runtime
2.
Consider reducing the buffer size. This reduces the number of variants annotated together in a batch and can be modified in both
command line and web interfaces. Reducing buffer size may increase run time.
3.
Ensure you are only using the options you need, rather than --everything. Some data-rich options, such as regulatory annotation
have an impact on memory use
!

Q: How to cite VEP?
If you use VEP, please cite our UPDATED publication so we can continue to support VEP development.
Web VEP questions
Q: How do I access the web version of the Variant Effect Predictor?
You can find the web VEP on the Tools page.
!
Q: Why is the output I get for my input file different when I use the web VEP and command line VEP?
Ensure that you are passing equivalent arguments to the script that you are using in the web version. If you are sure this is still a
problem, please report it on the ensembl-dev mailing list.
!
Q: Is there a tutorial for web VEP?
Yes, see our latest tutorial Annotating and prioritizing genomic variants using the Ensembl Variant Effect Predictor — A tutorial for more
information on using the Ensembl VEP web interface.
Command line VEP questions
Q: How can I make VEP run faster?
There are a number of factors that influence how fast VEP runs. Have a look at our handy guide for tips on improving VEP runtime.
!
Q: Why am I not seeing the same variant from my inout in the output?
Since the Ensembl 110 release, VEP by default will minimise the input allele for display in the output. To see the exact allele
representation you provided, use the --uploaded_allele option.
!
Q: Why do I see "N" as the reference allele in my HGVS strings?
Q: Why do I see the following error (or similar) in my VEP output?
substr outside of string at /nfs/users/nfs_w/wm2/Perl/ensembl-
variation/modules/Bio/EnsEMBL/Variation/Utils/Sequence.pm line 511.
Use of uninitialized value $ref_allele in string eq at /nfs/users/nfs_w/wm2/Perl/ensembl-
variation/modules/Bio/EnsEMBL/Variation/Utils/Sequence.pm line 514.
Use of uninitialized value in concatenation (.) or string at /nfs/users/nfs_w/wm2/Perl/ensembl-
variation/modules/Bio/EnsEMBL/Variation/Utils/Sequence.pm line 643.
Both of these error types are usually seen when using a FASTA file for retrieving sequence. There are a couple of steps you can take to
try to remedy them:
1.
The index alongside the FASTA can become corrupted. Delete [fastafile].index and re-run VEP to regenerate it. By default this file is
located in your $HOME/.vep/[species]/[version]_[assembly] directory.
2.
The FASTA file itself may have been corrupted during download; delete the fasta file and the index and re-download (you can use
the VEP installer to do this).
3.
Older versions of BioPerl (1.2.3 in particular is known to have this) cannot properly index large FASTA files. Make sure you are using
a later (>=1.6) version of BioPerl. The VEP installer installs 1.6.924 for you.
If you still see problems after taking these steps, or if you were not using a FASTA file in the first place, please contact us.
!
Q: Why do I see the following warning?

WARNING: Chromosome 21 not found in annotation sources or synonyms on line 160
This can occur if the chromosome names differ between your input variant and any annotation source that you are using (cache,
database, GFF/GTF file, FASTA file, custom annotation file). To circumvent this you may provide VEP with a synonyms file. A synonym
file is included in VEP's cache files, so if you have one of these for your species you can use it as follows:
./vep -i input.vcf -cache -synonyms ~/.vep/homo_sapiens/112_GRCh38/chr_synonyms.txt
The file consists of lines containing pairs of tab-separated synonyms. Order is not important as synonyms can be used in both
"directions".
!
Q: Can I get gnomAD exomes and genomes frequencies in VEP?
Yes, see this guide.
!
Q: Why do I see the following error?
Could not connect to database homo_sapiens_core_63_37 as user anonymous using
[DBI:mysql:database=homo_sapiens_core_63_37;host=ensembldb.ensembl.org;port=5306] as a locator:
Unknown MySQL server host 'ensembldb.ensembl.org' (2) at
$HOME/src/ensembl/modules/Bio/EnsEMBL/DBSQL/DBConnection.pm line 290.
-------------------- EXCEPTION --------------------
MSG: Could not connect to database homo_sapiens_core_63_37 as user anonymous using
[DBI:mysql:database=homo_sapiens_core_63_37;host=ensembldb.ensembl.org;port=5306] as a locator:
Unknown MySQL server host 'ensembldb.ensembl.org' (2)
By default VEP is configured to connect to the public MySQL server at ensembldb.ensembl.org. Occasionally the server may break
connection with your process, which causes this error. This can happen when the server is busy, or due to various network issues.
Consider using a local copy of the database, or the caching system.
!
Q: Can I use VEP on Windows?
Yes - see the documentation for a few different ways to get the VEP running on Windows.
!
Q: Can I use VEP with custom species and assemblies not available in Ensembl?
Yes - you can run VEP on any data you have by providing a custom GFF/GTF annotation and FASTA file, like so:
./vep -i input.vcf --gff data.gff.gz --fasta genome.fa.gz
!
Q: Can I download all of the SIFT and/or PolyPhen predictions?
The Ensembl Variation database and the human VEP cache file contain precalculated SIFT and PolyPhen-2 predictions for every
possible amino acid change in every translated protein product in Ensembl. Since these data are huge, we store them in a compressed
format. The best approach to extract them is to use our Perl API.
The format in which the data are stored in our database is described here
The simplest way to access these matrices is to use an API script to fetch a ProteinFunctionPredictionMatrix for your protein of interest
and then call its 'get_prediction' method to get the score for a particular position and amino acid, looping over all possible amino acids for
your position. There is some detailed documentation on this class in the API documentation here.
You would need to work out which peptide position your codon maps to, but there are methods in the TranscriptVariation class that
should help you (probably translation_start and translation_end).

!

Ensembl release 112 - January 2024 © EMBL-EBI
http://wp-np2-11..ebi.ac.uk
About Us
About us
Contact us
Citing Ensembl
Privacy policy
Disclaimer
Get help
Using this website
Adding custom tracks
Downloading data
Video tutorials
Variant Effect Predictor (VEP)
Our sister sites
Ensembl Bacteria
Ensembl Fungi
Ensembl Plants
Ensembl Protists
Ensembl Metazoa
Follow us
Blog
Twitter
Facebook
