Rsubread/Subread Users Guide
Rsubread v2.16.0/Subread v2.0.6
17 October 2023
Wei Shi and Yang Liao
Olivia Newton-John Cancer Research Institute
Melbourne, Australia
Copyright © 2011 - 2023
Contents
1 Introduction 3
2 Preliminaries 5
2.1 Citation . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 5
2.2 Download and installation . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 6
2.2.1 Install Bioconductor Rsubread package . . . . . . . . . . . . . . . . . . 6
2.2.2 Install SourceForge Subread package . . . . . . . . . . . . . . . . . . . . 6
2.3 How to get help . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 7
3 The seed-and-vote mapping paradigm 8
3.1 Seed-and-vote . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 8
3.2 Detection of short indels . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 9
3.3 Detection of exon-exon junctions . . . . . . . . . . . . . . . . . . . . . . . . . 10
3.4 Detection of structural variants (SVs) . . . . . . . . . . . . . . . . . . . . . . . 11
3.5 Two-scan read alignment . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 12
3.6 Multi-mapping reads . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 12
3.7 Mapping of paired-end reads . . . . . . . . . . . . . . . . . . . . . . . . . . . . 12
4 Mapping reads generated by genomic DNA sequencing technologies 14
4.1 A quick start for using SourceForge Subread package . . . . . . . . . . . . . . . 14
4.2 A quick start for using Bioconductor Rsubread package . . . . . . . . . . . . . 15
4.3 Index building . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 15
4.4 Read mapping . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 17
4.5 Memory use and speed . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 24
4.6 Mapping quality scores . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 24
4.7 Mapping output . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 24
4.8 Mapping of long reads . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 25
5 Mapping reads generated by RNA sequencing technologies 26
5.1 A quick start for using SourceForge Subread package . . . . . . . . . . . . . . . 26
5.2 A quick start for using Bioconductor Rsubread package . . . . . . . . . . . . . 27
5.3 Index building . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 28
5.4 Local read alignment . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 28
5.5 Global read alignment . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 28
1
5.6 Memory use and speed . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 29
5.7 Mapping output . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 29
5.8 Mapping microRNA sequencing reads (miRNA-seq) . . . . . . . . . . . . . . . 29
6 Read summarization 31
6.1 Introduction . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 31
6.2 featureCounts . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 32
6.2.1 Input data . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 32
6.2.2 Annotation format . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 32
6.2.3 In-built annotations . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 33
6.2.4 Single and paired-end reads . . . . . . . . . . . . . . . . . . . . . . . . 33
6.2.5 Assign reads to features and meta-features . . . . . . . . . . . . . . . . 34
6.2.6 Count multi-mapping reads and multi-overlapping reads . . . . . . . . 34
6.2.7 Read filtering . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 35
6.2.8 Read manipulation . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 36
6.2.9 Program output . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 36
6.2.10 Program usage . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 37
6.3 A quick start for featureCounts in SourceForge Subread . . . . . . . . . . . . . 45
6.4 A quick start for featureCounts in Bioconductor Rsubread . . . . . . . . . . . . 46
7 Quantify 10x scRNA-seq data 47
8 SNP calling 52
8.1 Algorithm . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 52
8.2 exactSNP . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 52
9 Utility programs 55
9.1 repair . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 55
9.2 flattenGTF . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 55
9.3 promoterRegions . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 55
9.4 propmapped . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 56
9.5 qualityScores . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 56
9.6 removeDup . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 56
9.7 subread-fullscan . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 56
9.8 txUnique . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 56
10 Case studies 57
10.1 A Bioconductor R pipeline for analyzing RNA-seq data . . . . . . . . . . . . . 57
2
Chapter 1
Introduction
The Subread/Rsubread packages comprise a suite of high-performance software programs
for processing next-generation sequencing data. Included in these packages are Subread
aligner, Subjunc aligner, Sublong long-read aligner, Subindel long indel detection program,
featureCounts read quantification program, exactSNP SNP calling program and other utility
programs. This document provides a detailed description to the programs included in the
packages.
Subread and Subjunc aligners adopt a mapping paradigm called “seed-and-vote” [1]. This
is an elegantly simple multi-seed strategy for mapping reads to a reference genome. This
strategy chooses the mapped genomic location for the read directly from the seeds. It uses a
relatively large number of short seeds (called subreads) extracted from each read and allows
all the seeds to vote on the optimal location. When the read length is <160 bp, overlapping
subreads are used. More conventional alignment algorithms are then used to fill in detailed
mismatch and indel information between the subreads that make up the winning voting block.
The strategy is fast because the overall genomic location has already been chosen before the
detailed alignment is done. It is sensitive because no individual subread is required to map
exactly, nor are individual subreads constrained to map close by other subreads. It is accurate
because the final location must be supported by several different subreads. The strategy
extends easily to find exon junctions, by locating reads that contain sets of subreads mapping
to different exons of the same gene. It scales up efficiently for longer reads.
Subread is a general-purpose read aligner. It can be used to align reads generated from
both genomic DNA sequencing and RNA sequencing technologies. It has been successfully
used in a number of high-profile studies [2, 3, 4, 5, 6]. Subjunc is specifically designed to detect
exon-exon junctions and to perform full alignments for RNA-seq reads. Note that Subread
performs local alignments for RNA-seq reads, whereas Subjunc performs global alignments for
RNA-seq reads. Subread and Subjunc comprise a read re-alignment step in which reads are
re-aligned using genomic variation data and junction data collected from the initial mapping.
The Subindel program carries out local read assembly to discover long insertions and
deletions. Read mapping should be performed before running this program.
The featureCounts program is designed to assign mapped reads or fragments (paired-end
data) to genomic features such as genes, exons and promoters. It is a light-weight read counting
3
program suitable for count both gDNA-seq and RNA-seq reads for genomic features[7]. The
Subread-featureCounts-limma/voom pipeline has been found to be one of the best-performing
pipelines for the analyses of RNA-seq data by the SEquencing Quality Control (SEQC) study,
the third stage of the well-known MicroArray Quality Control (MAQC) project [8].
Also included in this software suite is a very efficient SNP caller – ExactSNP. ExactSNP
measures local background noise for each candidate SNP and then uses that information to
accurately call SNPs.
These software programs support a variety of sequencing platforms. They are released in
two packages – SourceForge Subread package and Bioconductor Rsubread package[9].
4
Chapter 2
Preliminaries
2.1 Citation
If you use Rsubread, you can cite:
Liao Y, Smyth GK and Shi W (2019). The R package Rsubread is easier, faster,
cheaper and better for alignment and quantification of RNA sequencing reads.
Nucleic Acids Research, 47(8):e47.
http://www.ncbi.nlm.nih.gov/pubmed/30783653
If you use featureCounts, you can cite:
Liao Y, Smyth GK and Shi W (2014). featureCounts: an efficient general pur-
pose program for assigning sequence reads to genomic features. Bioinformatics,
30(7):923-30.
http://www.ncbi.nlm.nih.gov/pubmed/24227677
If you use Subread or Subjunc aligners, you can cite:
Liao Y, Smyth GK and Shi W (2013). The Subread aligner: fast, accurate and
scalable read mapping by seed-and-vote. Nucleic Acids Research, 41(10):e108.
http://www.ncbi.nlm.nih.gov/pubmed/23558742
If you use Rsubread inbuilt annotations, you can cite:
Chisanga D, Liao Y and Shi W (2022). Impact of gene annotation choice on the
quantification of RNA-seq data. BMC Bioinformatics, 23(1):107.
http://www.ncbi.nlm.nih.gov/pubmed/35354358
5
2.2 Download and installation
2.2.1 Install Bioconductor Rsubread package
R software needs to be installed on my computer before you can install this package. Launch
R and issue the following command to install Rsubread:
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install("Rsubread")
Alternatively you may download it from Rsubread web page http://bioconductor.org/
packages/release/bioc/html/Rsubread.html and install it manually.
2.2.2 Install SourceForge Subread package
Install from a binary distribution
This is the easiest way to install the SourceForge Subread package. Binary distributions are
available for Linux, Macintosh and Windows operating systems and they can be downloaded
from http://subread.sourceforge.net. The Linux binary distribution can be run on mul-
tiple Linux variants including Debian, Ubuntu, Fedora and Cent OS.
To install Subread package on FreeBSD or Solaris, you will have to install from source.
Install from source on a Unix or Macintosh computer
Download Subread source package to your working directory from SourceForge
http://subread.sourceforge.net, and type the following command to uncompress it:
tar zxvf subread-1.x.x.tar.gz
Enter src directory of the package and issue the following command to install it on a Linux
operating system:
make -f Makefile.Linux
To install it on a Mac OS X operating system, issue the following command:
make -f Makefile.MacOS
To install it on a FreeBSD operating system, issue the following command:
make -f Makefile.FreeBSD
6
To install it on Oracle Solaris or OpenSolaris computer operating systems, issue the fol-
lowing command:
make -f Makefile.SunOS
A new directory called bin will be created under the home directory of the software package,
and the executables generated from the compilation are saved to that directory. To enable
easy access to these executables, you may copy them to a system directory such as /usr/bin
or add the path to them to your search path (your search path is usually specified in the
environment variable ‘PATH’).
Install from source on a Windows computer
The MinGW software tool (http://www.mingw.org/) needs to installed to compile Subread.
2.3 How to get help
Bioconductor support site (https://support.bioconductor.org/) or Google Subread group
(https://groups.google.com/forum/#!forum/subread) are the best place to post questions
or make suggestions.
7

Chapter 3
The seed-and-vote mapping paradigm
3.1 Seed-and-vote
We have developed a new read mapping paradigm called “seed-and-vote” for efficient, accurate
and scalable read mapping [1]. The seed-and-vote strategy uses a number of overlapping seeds
from each read, called subreads. Instead of trying to pick the best seed, the strategy allows
all the seeds to vote on the optimal location for the read. The algorithm then uses more
conventional alignment algorithms to fill in detailed mismatch and indel information between
the subreads that make up the winning voting block. The following figure illustrates the
proposed seed-and-vote mapping approach with an toy example.
Two aligners have been developed under the seed-and-vote paradigm, including Subread
and Subjunc. Subread is a general-purpose read aligner, which can be used to map both
genomic DNA-seq and RNA-seq read data. Its running time is determined by the number of
subreads extracted from each read, not by the read length. Thus it has an excellent maping
scalability, ie. its running time has only very modest increase with the increase of read length.
8

Subread uses the largest mappable region in the read to determine its mapping location,
therefore it automatically determines whether a global alignment or a local alignment should
be found for the read. For the exon-spanning reads in a RNA-seq dataset, Subread performs
local alignments for them to find the target regions in the reference genome that have the
largest overlap with them. Note that Subread does not perform global alignments for the
exon-spanning reads and it soft clips those read bases which could not be mapped. However,
the Subread mapping result is sufficient for carrying out the gene-level expression analysis
using RNA-seq data, because the mapped read bases can be reliably used to assign reads,
including both exonic reads and exon-spanning reads, to genes.
To get the full alignments for exon-spanning RNA-seq reads, the Subjunc aligner can be
used. Subjunc is designd to discover exon-exon junctions from using RNA-seq data, but it
performs full alignments for all the reads at the same time. The Subjunc mapping results
should be used for detecting genomic variations in RNA-seq data, allele-specific expression
analysis and exon-level gene expression analysis. The Section 3.3 describes how exon-exon
junctions are discovered and how exon-spanning reads are aligned using the seed-and-vote
paradigm.
3.2 Detection of short indels
The seed-and-vote paradigm is very powerful in detecting short indels (insertions and
deletions). The figure below shows how we use the subreads to confidently detect short indels.
When there is an indel existing in a read, mapping locations of subreads extracted after the
indel will be shifted to the left (insertion) or to the right (deletion), relative to the mapping
9
locations of subreads at the left side of the indel. Therefore, indels in the reads can be
readily detected by examining the difference in mapping locations of the extracted subreads.
Moreover, the number of bases by which the mapping location of subreads are shifted gives the
precise length of the indel. Since no mismatches are allowed in the mapping of the subreads,
the indels can be detected with a very high accuracy.
3.3 Detection of exon-exon junctions
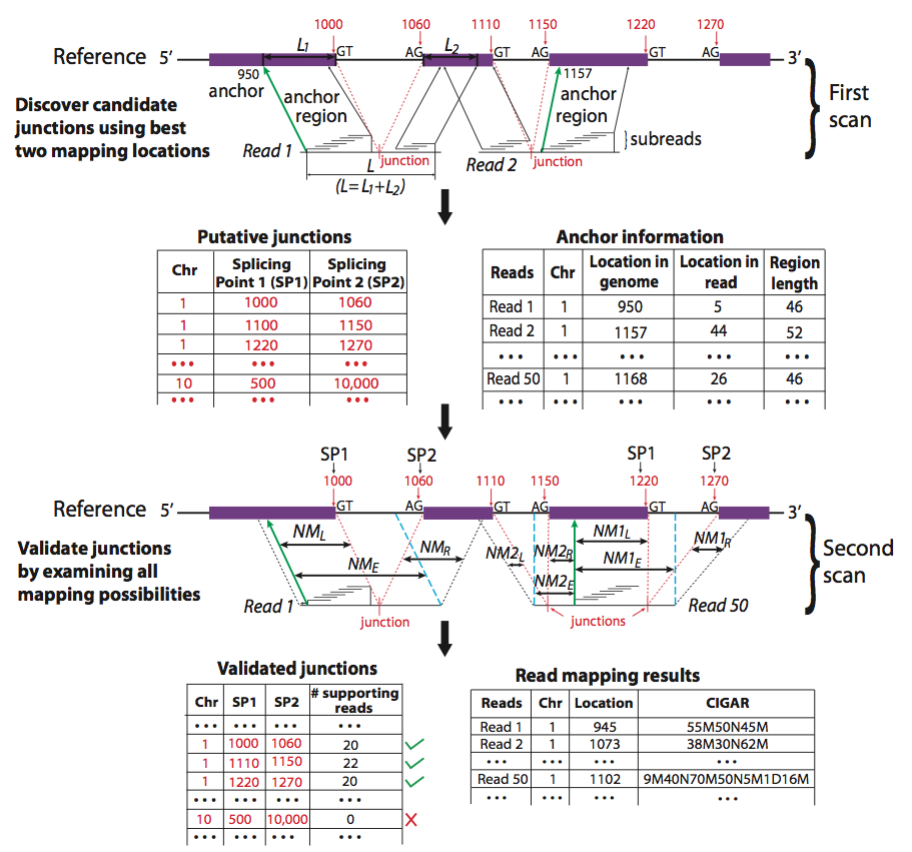
Figure below shows the schematic of exon-exon junction under seed-and-vote paradigm. The
first scan detects all possible exon-exon junctions using the mapping locations of the subreads
extracted from each read. Exons as short as 16bp can be detected in this step. The second scan
verifies the putative exon-exon junctions discovered from the first scan by read re-alignment.
This approach is implemented in the Subjunc program. The output of Subjunc includes
a list of discovered junctions, in addition to the mapping results. By default, Subjunc only
reports canonical exon-exon junctions that contain canonical donor and receptor sites (‘GT’
and ‘AG’ respectively). It was reported that such exon-exon junctions account for >98% of all
junctions. Orientation of donor and receptor sites is indicated by ‘XA’ tag in the SAM/BAM
output. Subjunc will report both canonical and non-canonical junctions when ‘–allJunctions’
option is turned on.
Accuracy of junction detection generally improves when external gene annotation data is
provided. The annotation data should include chromosomal coordinates of known exons of
each gene. Subjunc infers exon-exon junctions from the provided annotation data by connect-
ing each pair of neighboring exons from the same gene. This should cover majority of known
exon-exon junctions and the other junctions are expected to be discovered by the program.
Note that although Subread aligner does not report exon-exon junctions, providing this an-
notation is useful for it to map junction reads more accurately. See ‘-a’ parameter in Table 2
for more details.
10
3.4 Detection of structural variants (SVs)
Subread and Subjunc can be used detect SV events including long indel, duplication, inversion
and translocation, in RNA-seq and genomic DNA-seq data.
Detection of long indels is conducted by performing local read assembly. When the specified
indel length (‘-I’ option in SourceForge C or ‘indels’ paradigm in Rsubread) is greater than 16,
Subread and Subjunc will automatically start the read assembly process to detect long indels
(up to 200bp).
Breakpoints detected from SV events will be saved to a text file (‘.breakpoint.txt’), which
includes chromosomal coordinates of breakpoints and also the number of reads supporting
each pair of breakpoints found from the same SV event.
For the reads that were found to contain SV breakpoints, extra tags will be added for
11
them in mapping output. These tags include CC(chromosome name), CP(mapping position),
CG(CIGAR string) and CT(strand), and they describe the secondary alignment of the read
(the primary alignment is described in the main fields).
3.5 Two-scan read alignment
Subread and Subjunc aligners employ a two-scan approach for read mapping. In the first scan,
the aligners use seed-and-vote method to identify candidate mapping locations for each read
and also discover short indels, exon-exon junctions and structural variants. In the second
scan, they carry out final alignment for each read using the variant and junction information.
Variant and junction data (including chromosomal coordinates and number of supporting
reads) will be output along with the read mapping results. To the best of our knowledge,
Subread and Subjunc are the first to employ a two-scan mapping strategy to achieve a superior
mapping accuracy. This strategy was later seen in other aligners as well (called ‘two-pass’).
3.6 Multi-mapping reads
Multi-mapping reads are those reads that map to more than one genomic location with the
same similarity score (eg. number of mis-mismatched bases). Subread and Subjunc aligners
can effectively detect multi-mapping reads by closely examining candidate locations which
receive the highest number of votes or second highest number of votes. Numbers of mis-
matched bases and matched bases are counted for each candidate location during the final
re-alignment step and they are used for identifying multi-mapping reads. For RNA-seq data, a
read is called as a multi-mapping read if it has two or more candidate mapping locations that
have the same number of mis-matched bases and this number is the smallest in all candidate
locations being considered. For genomic DNA-seq data, a read is called as a multi-mapping
read if it has two or more candidate locations that have the same number of matched bases
and this number is the largest among all candidate locations being considered. Note that
for both RNA-seq and genomic DNA-seq data, any alignment reported for a multi-mapping
read must not have more than threshold number of mis-matched bases (as specified in ‘-M’
parameter).
For the reporting of a multi-mapping read, users may choose to not report any alignments
for the read (by default) or report up to a pre-defined number of alignments (‘–multiMapping’
and ‘-B’ options).
3.7 Mapping of paired-end reads
For the mapping of paired-end reads, we use the following formula to obtain a list of candidate
mapping locations for each read pair:
P E
score
= w ∗ (V
1
+ V
2
)
12
where V
1
and V
2
are the number of votes received from two reads from the same pair,
respectively. w has a value of 1.3 if mapping locations of the two reads are within the nominal
paired-end distance (or nominal fragment length), and has a value of 1 otherwise.
Up to 4,096 posssible alignments will be examined for each read pair and a maximum of
three candidate alignments with the highest P E
score
will be chosen for final re-alignment. Total
number of matched bases (for genomic DNA-seq data) or mis-matched bases (for RNA-seq
data) will be used to determine the best mapping in the final re-alignment step.
13

Chapter 4
Mapping reads generated by genomic
DNA sequencing technologies
4.1 A quick start for using SourceForge Subread package
An index must be built for the reference first and then the read mapping can be performed.
Step 1: Build an index
Build a base-space index (default). You can provide a list of FASTA files or a single FASTA
file including all the reference sequences. The files can be gzipped.
subread-buildindex -o my index chr1.fa chr2.fa ...
Step 2: Align reads
Map single-end genomic DNA sequencing reads using 5 threads (only uniquely mapped reads
are reported):
subread-align -t 1 -T 5 -i my index -r reads.txt.gz -o subread results.bam
Map paired-end reads:
subread-align -t 1 -d 50 -D 600 -i my index -r reads1.txt -R reads2.txt
-o subread results.bam
Detect indels of up to 16bp:
subread-align -t 1 -I 16 -i my index -r reads.txt -o subread results.bam
Report up to three best mapping locations:
subread-align -t 1 --multiMapping -B 3 -i my index -r reads.txt -o subread results.bam
14
4.2 A quick start for using Bioconductor Rsubread pack-
age
An index must be built for the reference first and then the read mapping can be performed.
Step 1: Building an index
To build the index, you must provide a single FASTA file (eg. “genome.fa”) which includes
all the reference sequences.
library(Rsubread)
buildindex(basename="my_index",reference="genome.fa")
Step 2: Aligning the reads
Map single-end reads using 5 threads:
align(index="my_index",readfile1="reads.txt.gz",type="dna",output_file="rsubread.bam",nthreads=5)
Detect indels of up to 16bp:
align(index="my_index",readfile1="reads.txt.gz",type="dna",output_file="rsubread.bam",indels=16)
Report up to three best mapping locations:
align(index="my_index",readfile1="reads.txt.gz",type="dna",output_file="rsubread.bam",
unique=FALSE,nBestLocations=3)
Map paired-end reads:
align(index="my_index",readfile1="reads1.txt.gz",readfile2="reads2.txt.gz",type="dna",
output_file="rsubread.bam",minFragLength=50,maxFragLength=600)
4.3 Index building
The subread-buildindex (buildindex function in Rsubread) program builds an index for refer-
ence genome by creating a hash table in which keys are 16bp mers (subreads) extracted from
the genome and values are their chromosomal locations.
A full index or a gapped index can be built for a reference genome. In a full index, subreads
are extracted from every location in the genome. In a gapped index, subreads are extracted
in every three bases in the genome (ie. there is a 2bp gap between two subreads next to each
other). When a full index is used in read mapping, only one set of subreads are extracted
from a read. However three sets of subreads need to be extracted from a read when a gapped
index is used for mapping. The first set starts from the first base of the read, the second set
starts from the second base and the third set starts from the third base. This makes sure that
a mapped read can always have a set of subreads that match those stored in the index.
15
A full index is larger than a gapped index. However the full index enables faster mapping
speed to be achieved. When a one-block full index is used for mapping, the maximum mapping
speed is achieved. Size of one-block full index built for the human reference genome (GRCh38)
is 17.8 GB. The subread-buildindex function needs 15 GB of memory to build this index.
Size of a gapped index built for GRCh38 is less than 9 GB and subread-buildindex needs 5.7
GB of memory to build it. Options are available to generate index of any size. In Rsubread,
a one-block full index is built by default.
The reference sequences should be in FASTA format. The subread-buildindex function
divides each reference sequence name (which can be found in the header lines) into multiple
substrings by using separators including ‘|’, ‘ ’(space) and ‘<tab>’, and it uses the first sub-
string as the name for the reference sequence during its index building. The first substrings
must be distinct for different reference sequences (otherwise the index cannot be built). Note
that the starting ‘>’ character in the header line is not included in the first substrings.
Sequences of reference genomes can be downloaded from public databases. For instance,
the primary assembly of human genome GRCh38 or mouse genome GRCm38 can be down-
loaded from the GENCODE database via the following links:
ftp://ftp.ebi.ac.uk/pub/databases/gencode/Gencode_human/release_28/GRCh38.primary_
assembly.genome.fa.gz
ftp://ftp.ebi.ac.uk/pub/databases/gencode/Gencode_mouse/release_M18/GRCm38.primary_
assembly.genome.fa.gz
Table 1 describes the arguments used by the subread-buildindex program.
16

Table 1: Arguments used by the subread-buildindex program (buildindex function in
Rsubread) in alphabetical order. Arguments in parenthesis in the first column are used by
buildindex.
Arguments Description
chr1.fa, chr2.fa, ...
(reference)
Give names of chromosome files. Note for Rsubread only a single
FASTA file including all reference sequences should be provided.
The files can be gzipped.
-B
(indexSplit=FALSE)
Create one block of index. The built index will not be split into
multiple pieces. The more blocks an index has, the slower the
mapping speed. This option will override ‘-M’ option when it is
also provided.
-c
(colorspace)
Build a color-space index.
-f < int >
(TH subread)
Specify the threshold for removing uninformative subreads (highly
repetitive 16bp mers). Subreads will be excluded from the index if
they occur more than threshold number of times in the reference
genome. Default value is 100.
-F
(gappedIndex=FALSE)
Build a full index for the reference genome. 16bp mers (subreads)
will be extracted from every position of a reference genome.
-M < int >
(memory)
Specify the size of computer memory(RAM) in megabytes that will
be used to store the index during read mapping, 8000MB by default.
If the index size is greater than the specified value, the index will
be split into multiple blocks. Only one block will be loaded into
memory at anytime during the read alignment.
-o < string >
(basename)
Specify the base name of the index to be created.
-v Output version of the program.
4.4 Read mapping
The Subread aligner (subread-align program in SourceForge Subread package or align func-
tion in Bioconductor Rsubread package) extracts a number of subreads from each read and
then uses these subreads to vote for the mapping location of the read. It uses the the “seed-
and-vote” paradigm for read mapping and reports the largest mappable region for each read.
Table 2 describes the arguments used by Subread aligner (also Subjunc and Sublong aligners).
Arguments used in Bioconductor Rsubread package are included in parenthesis.
17

Table 2: Arguments used by the subread-align/subjunc/sublong programs included in
the SourceForge Subread package in alphabetical order. Arguments in parenthesis in the
first column are the equivalent arguments used in Bioconductor Rsubread package.
(
1
subread-align arguments,
2
subjunc arguments and
3
sublong arguments)
Arguments Description
1,2
-a < string >
(useAnnotation,
annot.inbuilt, annot.ext)
Name of a gene annotation file that includes chromosomal
coordinates of exons from each gene. GTF/GFF format by
default. See -F option for supported formats. Users may
use the inbuilt annotations included in this package (SAF
format) for human and mouse data. Exon-exon junctions
are inferred by connecting each pair of neighboring exons
from the same gene. Gzipped file is accepted.
1,2
-A < string >
(chrAliases)
Name of a comma-delimited text file that includes aliases of
chromosome names. This file should contain two columns.
First column contains names of chromosomes included in
the SAF or GTF annotation and second column con-
tains corresponding names of chromosomes in the reference
genome. No column headers should be provided. Also note
that chromosome names are case sensitive. This file can be
used to match chromosome names between the annotation
and the reference genome.
1,2
-b
(color2base=TRUE)
Output base-space reads instead of color-space reads in
mapping output for color space data (eg. LifTech SOLiD
data). Note that the mapping itself will still be performed
at color-space.
1,2
-B < int >
(nBestLocations)
Specify the maximal number of equally-best mapping lo-
cations to be reported for a read. 1 by default. In the
mapping output, the ‘NH’ tag is used to indicate how
many alignments are reported for the read and the ‘HI’
tag is used for numbering the alignments reported for the
same read. This option should be used together with the
‘−−multiMapping’ option.
1,2
-d < int >
(minFragLength)
Specify the minimum fragment/template length, 50 by de-
fault. Note that if the two reads from the same pair do not
satisfy the fragment length criteria, they will be mapped
individually as if they were single-end reads.
1,2
-D < int >
(maxFragLength)
Specify the maximum fragment/template length, 600 by
default.
18
1,2
-F < string >
(isGTF)
Specify format of the provided annotation file. Acceptable
formats include ‘GTF’ (or compatible GFF format) and
‘SAF’. Default format in SourceForge Subread is ‘GTF’.
Default format in Rsubread is ‘SAF’.
1,2,3
-i < string >
(index)
Specify the base name of the index. The index used by
sublong aligner must be a full index and has only one block,
ie. ‘-F’ and ‘-B’ options must be specified when building
index with subread-buildindex.
1,2
-I < int >
(indels)
Specify the number of INDEL bases allowed in the map-
ping. 5 by default. Indels of up to 200bp long can be
detected.
1,2
-m < int >
(TH1)
Specify the consensus threshold, which is the minimal num-
ber of consensus subreads required for reporting a hit. The
consensus subreads are those subreads which vote for the
same location in the reference genome for the read. If pair-
end read data are provided, at least one of the two reads
from the same pair must satisfy this criteria. The default
value is 3 for subread-align, or 1 for subjunc and sublong.
1,2
-M < int >
(maxMismatches)
Specify the maximum number of mis-matched bases al-
lowed in the alignment. 3 by default. Mis-matches found
in soft-clipped bases are not counted.
1,2
-n < int >
(nsubreads)
Specify the number of subreads extracted from each read
for mapping. The default value is 10 for subread-align,
or 14 for subjunc. For sublong, this is number of subreads
(85 by default) extracted from each readlet. A readlet is a
100bp sequence extracted from a long read.
1,2,3
-o < string >
(output file)
Give the name of output file. The default output format is
BAM. All reads are included in mapping output, including
both mapped and unmapped reads, and they are in the
same order as in the input file.
1,2
-p < int >
(TH2)
Specify the minimum number of consensus subreads both
reads from the same pair must have. This argument is
only applicable for paired-end read data. The value of this
argument should not be greater than that of ‘-m’ option,
so as to rescue those read pairs in which one read has a
high mapping quality but the other does not. 1 by default.
1,2
-P < 3 : 6 >
(phredOffset)
Specify the format of Phred scores used in the input data,
’3’ for phred+33 and ’6’ for phred+64. ’3’ by default. For
align function in Rsubread, the possible values are ‘33’ (for
phred+33) and ‘64’ (for phred+64). ‘33’ by default.
19

1,2,3
-r < string >
(readfile1)
Give the name of an input file (multiple files are allowed to
be provided to align and subjunc functions in Rsubread).
For paired-end read data, this gives the first read file and
the other read file should be provided via the -R option.
Supported input formats include FASTQ/FASTA (uncom-
pressed or gzip compressed)(default), SAM and BAM.
1,2
-R < string >
(readfile2)
Provide name of the second read file from paired-end data.
The program will switch to paired-end read mapping mode
if this file is provided. (multiple files are allowed to be
provided to align and subjunc functions in Rsubread).
1,2
-S < ff : fr : rf >
(PE orientation)
Specify the orientation of the two reads from the same pair.
It has three possible values including ‘fr’, ‘ff’ and ‘’rf. Let-
ter ‘f’ denotes the forward strand and letter ‘r’ the reverse
strand. ‘fr’ by default (ie. the first read in the pair is on the
forward strand and the second read on the reverse strand).
1
-t < int >
(type)
Specify the type of input sequencing data. Possible values
include 0, denoting RNA-seq data, or 1, denoting genomic
DNA-seq data. User must specify the value. Character
values including ‘rna’ and ‘dna’ can also be used in the R
function. For genomic DNA-seq data, the aligner takes into
account both the number of matched bases and the num-
ber of mis-matched bases to determine the the best map-
ping location after applying the ‘seed-and-vote’ approach
for read mapping. For RNA-seq data, only the number of
mis-matched bases is considered for determining the best
mapping location.
1,2,3
-T < int >
(nthreads)
Specify the number of threads/CPUs used for mapping.
The value should be between 1 and 32. 1 by default.
20

2
−−allJunctions
(reportAllJunctions
=TRUE)
This option should be used with subjunc for de-
tecting canonical exon-exon junctions (with ‘GT/AG’
donor/receptor sites), non-canonical exon-exon junctions
and structural variants (SVs) in RNA-seq data. detected
junctions will be saved to a file with suffix name “.junc-
tion.bed”. Detected SV breakpoints will be saved to a
file with suffix name “.breakpoints.txt”, which includes
chromosomal coordinates of detected SV breakpoints and
also number of supporting reads. In the read map-
ping output, each breakpoint-containing read will contain
the following extra fields for the description of its sec-
ondary alignment: CC(Chr), CP(Position),CG(CIGAR)
and CT(strand). The primary alignment (described in the
main field) and secondary alignment give respectively the
mapping results for the two segments from the same read
that were seperated by the breakpoint. Note that each
breakpoint-containing read occupies only one row in map-
ping output. The mapping output includes mapping results
for all the reads.
1,2
−−BAMinput
(input format="BAM")
Specify that the input read data are in BAM format.
1,2
−−complexIndels Detect multiple short indels that occur concurrently in a
small genomic region (these indels could be as close as 1bp
apart).
1,2
−−DPGapExt < int >
(DP GapExtPenalty)
Specify the penalty for extending the gap when performing
the Smith-Waterman dynamic programming. 0 by default.
1,2
−−DPGapOpen < int >
(DP GapOpenPenalty)
Specify the penalty for opening a gap when applying the
Smith-Waterman dynamic programming to detecting in-
dels. -2 by default.
1,2
−−DPMismatch < int >
(DP MismatchPenalty)
Specify the penalty for mismatches when performing the
Smith-Waterman dynamic programming. 0 by default.
1,2
−−DPMatch < int >
(DP MatchScore)
Specify the score for the matched base when performing
the Smith-Waterman dynamic programming. 2 by default.
1,2
−−gtfFeature < string >
(GTF.featureType)
Specify the type of features that will be extracted from a
GTF annotation. ‘exon’ by default. Feature types can be
found in the 3rd column of a GTF annotation.
1,2
−−gtfAttr < string >
(GTF.attrType)
Specify the type of attributes in a GTF annotation that will
be used to group features. ‘gene id’ by default. Attributes
can be found in the 9th column of a GTF annotation.
21

1,2
−−keepReadOrder
(keepReadOrder
=FALSE)
Output reads in the same order as in the input read file.
This option only applies to BAM output. Note that in the
output, reads from the same pair are always placed next to
each other no matter this option is provided or not.
1,2
−−multiMapping
(unique=FALSE)
Multi-mapping reads will also be reported in the mapping
output. Number of alignments reported for each multi-
mapping read is determined by the ‘-B’ option. If the total
number of equally best mapping locations found for a read
is greater than the number specified by ‘-B’, then random
mapping locations (total number of these locations is the
same as ‘-B’ value) will be selected. For example, if value
of ‘-B’ is 1, then one random location will be reported.
1,2
−−rg < string >
(readGroup)
Add a < tag : value > to the read group (RG) header in
the mapping output.
1,2
−−rg-id < string >
(readGroupID)
Specify the read group ID. If specified, the read group ID
will be added to the read group header field and also to
each read in the mapping output.
1,2
−−SAMinput
(input format="SAM")
Specify that the input read data are in SAM format.
1,2,3
−−SAMoutput
(output format="SAM")
Specify that mapping results are saved into a SAM format
file.
1,2
−−sortReadsByCoordinates
(sortReadsByCoordinates
=FALSE)
If specified, reads will be sorted by their mapping coordi-
nates in the mapping output. This option is applicable for
BAM output only. A BAI index file will also be generated
for each BAM file so the BAM files can be directly loaded
into a genome browser such as IGB or IGV.
1
−−sv
(detectSV=TRUE)
This option should be used with subread-align for detect-
ing structural variants (SVs) in genomic DNA sequencing
data. Detected SV breakpoints will be saved to a file with
suffix name “.breakpoints.txt”, which includes chromoso-
mal coordinates of detected SV breakpoints and also num-
ber of supporting reads for each SV event. In the read
mapping output, each breakpoint-containing read will con-
tain the following extra fields for the description of its sec-
ondary alignment: CC(Chr), CP(Position),CG(CIGAR)
and CT(strand). The primary alignment (described in the
main field) and secondary alignment give respectively the
mapping results for the two segments from the same read
that were seperated by the breakpoint. Note that each
breakpoint-containing read occupies only one row in map-
ping output. The mapping output includes mapping results
for all the reads.
22

1,2
−−trim5 < int >
(nTrim5)
Trim off < int > number of bases from 5’ end of each read.
0 by default.
1,2
−−trim3 < int >
(nTrim3)
Trim off < int > number of bases from 3’ end of each read.
0 by default.
1,2,3
-v Output version of the program.
23

4.5 Memory use and speed
subread-buildindex (buildindex function in Rsubread) needs 15GB of memory to build a
full index for human/mouse genome. With this index, subread-align (align in Rsubread)
require 17.8GB of memory for read mapping. This enables fastest mapping speed, but it is
recommended that the full index should be on a unix server due to relatively large memory
use. Mapping rate is ∼14 million reads per minute (10 CPU threads) when full index is used.
A gapped index is recommended for use on a personal computer, which typically has
16GB of memory or less. subread-buildindex (buildindex function in Rsubread) only needs
5.7GB of memory to build a gapped index for human/mouse genome. subread-align (align
in Rsubread) needs 8.2GB of memory for mapping with the gapped index.
It takes subread-buildindex (buildindex function in Rsubread) about 40 minutes to build
a full index for human/mouse genome, and building a gapped index takes about 15 minutes.
Memory use for index building and read mapping can be further reduced by building a
split index using the -B and -M options in subread-buildindex (indexSplit and memory options
in buildindex function in Rsubread).
4.6 Mapping quality scores
Subread and Subjunc aligners determine the final mapping location of each read by taking into
account vote number, number of mis-matched bases, number of matched bases and mapping
distance between two reads from the same pair (for paired-end reads only) . They then assign
a mapping quality score (MQS) to each mapped read to indicate the confidence of mapping
using the following formula:
MQS =
40
(N
c
+N
mm
)
if only one best location found
0 if > 1 equally best locations were found
where N
c
is the number of candidate locations considered at the re-alignment step (note that
no more than three candidate locations are considered at this step). N
mm
is the number of
mismatches present in the final reported alignment for the read.
4.7 Mapping output
Read mapping results for each library will be saved to a BAM or SAM format file. Short indels
detected from the read data will be saved to a text file (‘.indel’). If ‘−−sv’ is specified when
running subread-align, breakpoints detected from structural variant events will be output to
a text file for each library as well (‘.breakpoints.txt’). Screen output includes a brief mapping
24
summary, including percentage of uniquely mapped reads, percentage of multi-mapping reads
and percentage of unmapped reads.
4.8 Mapping of long reads
We developed a new long-read aligner, called Sublong, for the mapping of long reads that were
generated by long-read sequencing technologies such as Nanopore and PacBio sequencers.
Sublong is also based on the seed-and-vote mapping strategy. Parameters of Sublong program
can be found in Table 2.
25

Chapter 5
Mapping reads generated by RNA
sequencing technologies
5.1 A quick start for using SourceForge Subread package
An index must be built for the reference first and then the read mapping and/or junction
detection can be carried out.
Step 1: Building an index
The following command can be used to build a base-space index. You can provide a list of
FASTA files or a single FASTA file including all the reference sequences. The files can be
gzipped.
subread-buildindex -o my index chr1.fa chr2.fa ...
For more details about index building, see Section 4.3.
Step 2: Aligning the reads
Subread
If the purpose of an RNA-seq experiment is to quantify gene-level expression and discover
differentially expressed genes, the Subread aligner is recommended. Subread carries out local
alignments for RNA-seq reads. The commands used by Subread to align RNA-seq reads are
the same as those used to align gDNA-seq reads. Below is an example of using Subread to
map single-end RNA-seq reads.
subread-align -t 0 -i my index -r rnaseq-reads.txt -o subread results.bam
Another RNA-seq aligner included in this package is the Subjunc aligner. Subjunc not only
26

performs read alignments but also detects exon-exon junctions. The main difference between
Subread and Subjunc is that Subread does not attempt to detect exon-exon junctions in the
RNA-seq reads. For the alignments of the exon-spanning reads, Subread just uses the largest
mappable regions in the reads to find their mapping locations. This makes Subread more
computationally efficient. The largest mappable regions can then be used to reliably assign
the reads to their target genes by using a read summarization program (eg. featureCounts, see
Section 6.2), and differential expression analysis can be readily performed based on the read
counts yielded from read summarization. Therefore, Subread is sufficient for read mapping if
the purpose of RNA-seq analysis is to perform a differential expression analysis. Also, Subread
could report more mapped reads than Subjunc. For example, the exon-spanning reads that
are not aligned by Subjunc due to the lack of canonical GT/AG splicing signals can be aligned
by Subread as long as they have a good match with the reference sequence.
Subjunc
For other purposes of the RNA-seq data anlayses such as exon-exon junction detection,
alternative splicing analysis and genomic mutation detection, Subjunc aligner should be used
because exon-spanning reads need to be fully aligned. Below is an example command of using
Subjunc to perform global alignments for paired-end RNA-seq reads. Note that there are two
files produced after mapping: one is a BAM-format file including mapping results and the
other a BED-format file including discovered exon-exon junctions.
subjunc -i my index -r rnaseq-reads1.txt -R rnaseq-reads2.txt -o subjunc result
5.2 A quick start for using Bioconductor Rsubread pack-
age
An index must be built for the reference first and then the read mapping can be performed.
Step 1: Building an index
To build the index, you must provide a single FASTA file (eg. “genome.fa”) which includes
all the reference sequences. The FASTA file can be a gzipped file.
library(Rsubread)
buildindex(basename="my_index",reference="genome.fa")
Step 2: Aligning the reads
Please refer to Section 5.1 for difference between Subread and Subjunc in mapping RNA-
seq data. Below is an example for mapping a single-end RNA-seq dataset using Subread.
Useful information about align function can be found in its help page (type ?align in your
R prompt).
27
align(index="my_index",readfile1="rnaseq-reads.txt.gz",output_file="subread_results.bam")
Below is an example for mapping a single-end RNA-seq dataset using Subjunc. Useful
information about subjunc function can be found in its help page (type ?subjunc in your R
prompt).
subjunc(index="my_index",readfile1="rnaseq-reads.txt.gz",output_file="subjunc_results.bam")
5.3 Index building
Please refer to Section 4.3. Same index is used for the mapping of RNA and DNA sequencing
reads.
5.4 Local read alignment
The Subread and Subjunc can both be used to map RNA-seq reads to the reference genome.
If the goal of the RNA-seq data is to perform expression analysis, eg. finding genes expressing
differentially between different conditions, then Subread is recommended. Subread performs
fast local alignments for reads and reports the mapping locations that have the largest overlap
with the reads. These reads can then be assigned to genes for expression analysis. For this
type of analysis, global alignments for the exon-spanning reads are not required because local
aligments are sufficient to get reads to be accurately assigned to genes.
However, for other types of RNA-seq data analyses such as exon-exon junction discovery,
genomic mutation detection and allele-specific gene expression analysis, global alignments are
required. The next section describes the Subjunc aligner, which performs global aligments for
RNA-seq reads.
5.5 Global read alignment
Subjunc aligns each exon-spanning read by firstly using a large number of subreads extracted
from the read to identify multiple target regions matching the selected subreads, and then
using the splicing signals (donor and receptor sites) to precisely determine the mapping loca-
tions of the read bases. It also includes a verification step to compare the quality of mapping
reads as exon-spanning reads with the quality of mapping reads as exonic reads to finally
decide how to best map the reads. Reads may be re-aligned if required.
Output of Subjunc aligner includes a list of discovered exon-exon junction locations and
also the complete alignment results for the reads. Table 2 describes the arguments used by
the Subjunc program.
28

5.6 Memory use and speed
Memory use and running time of subread-buildindex and subread-align (buildindex and
align in Rsubread) are the same as their memory use and running time in the analysis of DNA
sequencing data (see Section 4.5).
Compared to subread-align (align in Rsubread), subjunc uses the same amount of memory
when a full index is used and it uses slightly more memory (8.8GB of memory for human/mouse
data) when a gapped index is used. subjunc is also slightly slower than subread-align.
5.7 Mapping output
Read mapping results for each library will be saved to a BAM/SAM file. Detected exon-exon
junctions will be saved to a BED file for each library (‘.junction.bed’). Detected short indels
will be saved to a text file (‘.indel’). Screen output includes a brief mapping summary, includ-
ing percentage of uniquely mapped reads, percentage of multi-mapping reads and percentage
of unmapped reads.
5.8 Mapping microRNA sequencing reads (miRNA-seq)
To use Subread aligner to map miRNA-seq reads, a full index must be built for the reference
genome before read mapping can be carried out. For example, the following command builds
a full index for mouse reference genome mm39 :
subread-buildindex -F -B -o mm39 full index mm39.fa
The full index includes 16bp mers extracted from every genomic location in the genome.
Note that if -F is not specified, subread-buildindex builds a gapped index which includes
16bp mers extracted every three bases in the reference genome, ie. there is a 2bp gap between
each pair of neighbouring 16bp mers.
After the full index was built, read alignment can be performed. Reads do not need to be
trimmed before feeding them to Subread aligner since Subread soft clips sequences in the reads
that can not be properly mapped. The parameters used for mapping miRNA-seq reads need
to be carefully designed due to the very short length of miRNA sequences (∼22bp). The total
number of subreads (16bp mers) extracted from each read should be the read length minus
15, which is the maximum number of subreads that can be possibly extracted from a read.
The reason why we need to extract the maximum number of subreads is to achieve a high
sensitivity in detecting the short miRNA sequences.
The threshold for the number of consensus subreads required for reporting a hit should be
within the range of 2 to 7 consensus subreads inclusive. The larger the number of consensus
subreads required, the more stringent the mapping will be. Using a threshold of 2 consensus
subreads allows the detection of miRNA sequences of as short as 17bp, but the mapping error
rate could be relatively high. With this threshold, there will be at least 17 perfectly matched
29

bases present in each reported alignment. If a threshold of 4 consensus subreads was used,
length of miRNA sequences that can be detected is 19 bp or longer. With this threshold,
there will be at least 19 perfectly matched bases present in each reported alignment. When a
threshold of 7 consensus subreads was used, only miRNA sequences of 22bp or longer can be
detected (at least 22 perfectly matched bases will be present in each reported alignment).
We found that there was a significant decrease in the number of mapped reads when the
requried number of consensus subreads increased from 4 to 5 when we tried to align a mouse
miRNA-seq dataset, suggesting that there are a lot of miRNA sequences that are only 19bp
long. We therefore used a threshold of 4 consensus subreads to map this dataset. However,
what we observed might not be the case for other datasets that were generated from different
cell types and different species.
Below is an example of mapping 50bp long reads (adaptor sequences were included in the
reads in addition to the miRNA sequences), with at least 4 consensus subreads required in
the mapping. Note that ‘-t’ option should have a value of 1 since miRNA-seq reads are more
similar to gDNA-seq reads than mRNA-seq reads from the read mapping point of vew.
subread-align -t 1 -i mm39 full index -n 35 -m 4 -M 3 -T 10 -I 0 --multiMapping -B 10
-r miRNA reads.fastq -o result.bam
The ‘-B 10’ parameter instructs Subread aligner to report up to 10 best mapping locations
(equally best) in the mapping results. The multiple locations reported for the reads could
be useful for investigating their true origin, but they might need to be filtered out when
assigning mapped reads to known miRNA genes to ensure a high-quality quantification of
miRNA genes. The miRBase database (http://www.mirbase.org/) is a useful resource that
includes annotations for miRNA genes in many species. The featureCounts program can be
readily used for summarizing reads to miRNA genes.
30
Chapter 6
Read summarization
6.1 Introduction
Sequencing reads often need to be assigned to genomic features of interest after they are
mapped to the reference genome. This process is often called read summarization or read
quantification. Read summarization is required by a number of downstream analyses such as
gene expression analysis and histone modification analysis. The output of read summarization
is a count table, in which the number of reads assigned to each feature in each library is
recorded.
A particular challenge to the read summarization is how to deal with those reads that
overlap more than one feature (eg. an exon) or meta-feature (eg. a gene). Care must be
taken to ensure that such reads are not over-counted or under-counted. Here we describe
the featureCounts program, an efficient and accurate read quantifier. featureCounts has the
following features:
• It carries out precise and accurate read assignments by taking care of indels, junctions
and structural variants in the reads.
• It takes only half a minute to summarize 20 million reads.
• It supports GTF and SAF format annotation.
• It supports strand-specific read counting.
• It can count reads at feature (eg. exon) or meta-feature (eg. gene) level.
• Highly flexible in counting multi-mapping and multi-overlapping reads. Such reads can
be excluded, fully counted or fractionally counted.
• It gives users full control on the summarization of paired-end reads, including allowing
them to check if both ends are mapped and/or if the fragment length falls within the
specified range.
31
• Reduce ambiguity in assigning read pairs by searching features that overlap with both
reads from the pair.
• It allows users to specify whether chimeric fragments should be counted.
• Automatically detect input format (SAM or BAM).
• Automatically sort paired-end reads. Users can provide either location-sorted or name-
sorted bams files to featureCounts. Read sorting is implemented on the fly and it only
incurs minimal time cost.
6.2 featureCounts
6.2.1 Input data
The data input to featureCounts consists of (i) one or more files of aligned reads (short or
long reads) in either SAM or BAM format and (ii) a list of genomic features in either Gene
Transfer Format (GTF) or General Feature Format (GFF) or Simplified Annotation Format
(SAF). The format of input reads is automatically detected (SAM or BAM).
If the input contains location-sorted paired-end reads, featureCounts will automatically
re-order the reads to place next to each other the reads from the same pair before counting
them. Sometimes name-sorted paired-end input reads are not compatible with featureCounts
(due to for example reporting of multi-mapping results) and in this case featureCounts will
also automatically re-order them. We provide an utility program repair to allow users to pair
up the reads before feeding them to featureCounts.
Both read alignment and read counting should use the same reference genome. For each
read, the BAM/SAM file gives the name of the reference chromosome or contig the read
mapped to, the start position of the read on the chromosome or contig/scaffold, and the
so-called CIGAR string giving the detailed alignment information including insertions and
deletions and so on relative to the start position.
The genomic features can be specified in either GTF/GFF or SAF format. The SAF format
is the simpler and includes only five required columns for each feature (see next section). In
either format, the feature identifiers are assumed to be unique, in accordance with commonly
used Gene Transfer Format (GTF) refinement of GFF.
featureCounts supports strand-specific read counting if strand-specific information is pro-
vided. Read mapping results usually include mapping quality scores for mapped reads. Users
can optionally specify a minimum mapping quality score that the assigned reads must satisfy.
6.2.2 Annotation format
The genomic features can be specified in either GTF/GFF or SAF format. A definition of the
GTF format can be found at UCSC website (http://genome.ucsc.edu/FAQ/FAQformat.
html#format4). The SAF format includes five required columns for each feature: feature
identifier, chromosome name, start position, end position and strand. Both start and end
32
positions are inclusive. These five columns provide the minimal sufficient information for
read quantification purposes. Extra annotation data are allowed to be added from the sixth
column.
A SAF-format annotation file should be a tab-delimited text file. It should also include a
header line. An example of a SAF annotation is shown as below:
GeneID Chr Start End Strand
497097 chr1 3204563 3207049 -
497097 chr1 3411783 3411982 -
497097 chr1 3660633 3661579 -
100503874 chr1 3637390 3640590 -
100503874 chr1 3648928 3648985 -
100038431 chr1 3670236 3671869 -
...
GeneID column includes gene identifiers that can be numbers or character strings. Chro-
mosomal names included in the Chr column must match the chromosomal names of reference
sequences to which the reads were aligned.
6.2.3 In-built annotations
In-built gene annotations for genomes hg38, hg19, mm39, mm10 and mm9 are included in
both Bioconductor Rsubread package and SourceForge Subread package. These annotations
were downloaded from NCBI RefSeq database and then adapted by merging overlapping exons
from the same gene to form a set of disjoint exons for each gene. Genes with the same Entrez
gene identifiers were also merged into one gene.
Each row in the annotation represents an exon of a gene. There are five columns in
the annotation data including Entrez gene identifier (GeneID), chromosomal name (Chr ),
chromosomal start position(Start), chromosomal end position (End) and strand (Strand).
In Rsubread, users can access these annotations via the getInBuiltAnnotation function. In
Subread, these annotations are stored in directory ‘annotation’ under home directory of the
package.
6.2.4 Single and paired-end reads
Reads may be paired or unpaired. If paired reads are used, then each pair of reads defines
a DNA or RNA fragment bookended by the two reads. In this case, featureCounts can be
instructed to count fragments rather than reads. featureCounts automatically sorts reads by
name if paired reads are not in consecutive positions in the SAM or BAM file, with minimal
cost. Users do not need to sort their paired reads before providing them to featureCounts.
33
6.2.5 Assign reads to features and meta-features
featureCounts is a general-purpose read summarization function, which assigns mapped reads
(RNA-seq reads or genomic DNA-seq reads) to genomic features or meta-features. A feature
is an interval (range of positions) on one of the reference sequences. A meta-feature is a
set of features that represents a biological construct of interest. For example, features often
correspond to exons and meta-features to genes. Features sharing the same feature identifier
in the GTF or SAF annotation are taken to belong to the same meta-feature. featureCounts
can summarize reads at either the feature or meta-feature levels.
We recommend to use unique gene identifiers, such as NCBI Entrez gene identifiers, to
cluster features into meta-features. Gene names are not recommended to use for this purpose
because different genes may have the same names. Unique gene identifiers were often included
in many publicly available GTF annotations which can be readily used for summarization.
The Bioconductor Rsubread package also includes NCBI RefSeq annotations for human and
mice. Entrez gene identifiers are used in these annotations.
featureCounts preforms precise read assignment by comparing mapping location of every
base in the read with the genomic region spanned by each feature. It takes account of any
gaps (insertions, deletions, exon-exon junctions or structural variants) that are found in the
read. It calls a hit if any overlap is found between read and feature.
Users may use ‘–minOverlap (minOverlap in R)’ and ‘–fracOverlap (fracOverlap in R)’
options to specify the minimum number of overlapping bases and minimum fraction of over-
lapping bases requried for assigning a read to a feature, respectively. The ‘–fracOverlap’ option
might be particularly useful for counting reads with variable lengths.
When counting reads at meta-feature level, a hit is called for a meta-feature if the read
overlaps any component feature of the meta-feature. Note that if a read hits a meta-feature,
it is always counted once no matter how many features in the meta-feature this read overalps
with. For instance, an exon-spanning read overlapping with more than one exon within the
same gene only contributes 1 count to the gene.
When assigning reads to genes or exons, most reads can be successfully assigned without
ambiguity. However if reads are to be assigned to transcripts, due to the high overlap between
transcripts from the same gene, many reads will be found to overlap more than one transcript
and therefore cannot be uniquely assigned. Specialized transcript-level quantification tools
are recommended for counting reads to transcripts. Such tools use model-based approaches
to deconvolve reads overlapping with multiple transcripts.
6.2.6 Count multi-mapping reads and multi-overlapping reads
A multi-mapping read is a read that maps to more than one location in the reference genome.
There are multiple options for counting such reads. Users can specify the ‘-M’ option (set
countMultiMappingReads to TRUE in R) to fully count every alignment reported for a multi-
mapping read (each alignment carries 1 count), or specify both ‘-M’ and ‘–fraction’ options (set
both countMultiMappingReads and fraction to TRUE in R) to count each alignment fractionally
(each alignment carries 1/x count where x is the total number of alignments reported for the
34
read), or do not count such reads at all (this is the default behavior in SourceForge Subread
package; In R, you need to set countMultiMappingReads to FALSE).
A multi-overlapping read is a read that overlaps more than one meta-feature when counting
reads at meta-feature level or overlaps more than one feature when counting reads at feature
level. The decision of whether or not to counting these reads is often determined by the
experiment type. We recommend that reads or fragments overlapping more than one gene
are not counted for RNA-seq experiments, because any single fragment must originate from
only one of the target genes but the identity of the true target gene cannot be confidently
determined. On the other hand, we recommend that multi-overlapping reads or fragments are
counted for ChIP-seq experiments because for example epigenetic modifications inferred from
these reads may regulate the biological functions of all their overlapping genes.
By default, featureCounts does not count multi-overlapping reads. Users can specify the
‘-O’ option (set allowMultiOverlap to TRUE in R) to fully count them for each overlapping meta-
feature/feature (each overlapping meta-feature/feature receives a count of 1 from a read), or
specify both ‘-O’ and ‘–fraction’ options (set both allowMultiOverlap and fraction to TRUE
in R) to assign a fractional count to each overlapping meta-feature/feature (each overlapping
meta-feature/feature receives a count of 1/y from a read where y is the total number of
meta-features/features overlapping with the read).
If a read is both multi-mapping and multi-overlapping, then when ‘-O’, ‘-M’, and ‘–fraction’
are all specified each overlapping meta-feature/feature will receive a fractional count of 1/(x ∗
y). Note that each alignment reported for a multi-mapping read is assessed separately for
overlapping with multiple meta-features/features.
When multi-mapping reads are reported with primary and secondary alignments and both
‘-M’ and ‘–primary’ are specified, only primary alignments will be considered in counting and
secondary alignments will be ignored. If ‘-M’ is specified but ‘–primary’ is not specified, both
primary and secondary alignments will be considered in counting. Note that all the alignments
reported for a multi-mapping read are expected to have a ‘NH’ tag and whether an alignment
is primary or secondary is determined by using bit 0x100 in the FLAG field of the alignment
record.
6.2.7 Read filtering
featureCounts implements a variety of read filters to facilitate flexible read counting, which
should satisfy the requirement of most downstream analyses. The order of these filters being
applied is as following (from first to last): unmapped > read type > singleton > mapping
quality > chimeric fragment > fragment length > duplicate > multi-mapping > secondary
alignment > split reads (or nonsplit reads) > no overlapping features > overlapping length >
assignment ambiguity.
Number of reads that were excluded from counting by each filter is reported in the program
output, in addition to the reported read counts (see Section 6.2.9). The ‘read type’ filter
removes those reads that have an unexpected read type and also cannot be counted with
confidence. For example, if there are single end reads included in a paired end read dataset
(such data can be produced from a read trimming program for instance) and reads are required
35

to be counted in a strand-specific manner, then all the single end reads will be excluded from
counting because their strandness cannot be determined. However if such reads are to be
counted in an unstranded manner then all the single end reads will be considered for counting.
6.2.8 Read manipulation
Reads can be shifted (--readShiftType and --readShiftSize), extended (--readExtension5
and --readExtension3) or reduced to an end base (--read2pos), before being assigned to
features/meta-features. These read manipulations are carried out by featureCounts in the
following order: shift > extension > reduction.
6.2.9 Program output
The output of featureCounts program includes a count table and a summary of counting results.
For SourceForge Subread, the output data are saved to two tab-delimited files: one file contains
read counts (file name is specified by the user) and the other file includes summary of counting
results (file name is the name of read count file added with ‘.summary’). For Rsubread, all the
output data are saved to an R ‘List’ object (for more details see the help page for featureCounts
function in Rsubread package).
The read count table includes annotation columns (‘Geneid’, ‘Chr’, ‘Start’, ‘End’, ‘Strand’
and ‘Length’) and data columns (eg. read counts for genes for each library). When counting
reads to meta-features (eg. genes) columns ‘Chr’, ‘Start’, ‘End’ and ‘Strand’ may each contain
multiple values (separated by semi-colons), which correspond to individual features included
in the same meta-feature. Column ‘Length’ always contains one single value which is the
total number of non-overlapping bases included in a meta-feature (or a feature), regardless
of counting at meta-feature level or feature level. When counting RNA-seq reads to genes,
the ‘Length’ column typically contains the total number of non-overlapping bases in exons
belonging to the same gene for each gene.
The counting summary includes total number of alignments that were successfully assigned
and also number of alignments that failed to be assigned due to various filters. Note that the
counting summary includes the number of alignments, not the number of reads. Number of
alignments will be higher than the number of reads when multi-mapping reads are included
since each multi-mapping read contains more than one alignment. Number and percentage of
successfully assigned alignments are also shown in featureCounts screen output.
Filters supported by featureCounts can be found in the list below:
• Unassigned Unmapped: unmapped reads cannot be assigned.
• Unassigned Read Type: reads that have an unexpected read type (eg. being a single
end read included in a paired end dataset) and also cannot be counted with confidence
(eg. due to stranded counting). Such reads are typically generated from a read trimming
program.
• Unassigned Singleton: read pairs that have only one end mapped.
36

• Unassigned MappingQuality: alignments with a mapping quality score lower than the
threshold.
• Unassigned Chimera: two ends in a paired end alignment are located on different chro-
mosomes or have unexpected orientation.
• Unassigned FragementLength: fragment length inferred from paired end alignment does
not meet the length criteria.
• Unassigned Duplicate: alignments marked as duplicate (indicated in the FLAG field).
• Unassigned MultiMapping: alignments reported for multi-mapping reads (indicated by
‘NH’ tag).
• Unassigned Secondary: alignments reported as secondary alignments (indicated in the
FLAG field).
• Unassigned Split (or Unassigned NonSplit): alignments that contain junctions (or do
not contain junctions).
• Unassigned NoFeatures: alignments that do not overlap any feature.
• Unassigned Overlapping Length: alignments that do not overlap any feature (or meta-
feature) with the minimum required overlap length.
• Unassigned Ambiguity: alignments that overlap two or more features (feature-level sum-
marization) or meta-features (meta-feature-level summarization).
In the counting summary these filters are listed in the same order as they were applied
in counting process (see Section 6.2.7). An unassigned alignment might fall into more than
one category as listed above, however it will only be allocated to one category which is the
category corresponding to the first filter that filtered this alignment out.
6.2.10 Program usage
Table 3 describes the parameters used by the featureCounts program.
37

Table 3: Arguments used by the featureCounts program included in the SourceForge Sub-
read package in alphabetical order. Arguments included in parenthesis are the equivalent
parameters used by featureCounts function in Bioconductor Rsubread package.
Arguments Description
input files
(files)
Give the names of input read files that include the read map-
ping results. The program automatically detects the file for-
mat (SAM or BAM). Multiple files can be provided at the
same time. Files are allowed to be provided via < stdin >.
-a < string >
(annot.ext,
annot.inbuilt)
Provide name of an annotation file. See -F option for file
format. Gzipped file is accepted.
-A
(chrAliases)
Provide a chromosome name alias file to match chr names in
annotation with those in the reads. This should be a two-
column comma-delimited text file. Its first column should
include chr names in the annotation and its second column
should include chr names in the reads. Chr names are case
sensitive. No column header should be included in the file.
-B
(requireBothEndsMapped)
If specified, only fragments that have both ends successfully
aligned will be considered for summarization. This option is
only applicable for the counting of fragments (read pairs).
-C
(countChimericFragments)
If specified, the chimeric fragments (those fragments that have
their two ends aligned to different chromosomes) will NOT be
counted. This option is only applicable for the counting of
fragments (read pairs).
-d < int >
(minFragLength)
Minimum fragment/template length, 50 by default.
-D < int >
(maxFragLength)
Maximum fragment/template length, 600 by default.
-f
(useMetaFeatures)
If specified, read summarization will be performed at feature
level (eg. exon level). Otherwise, it is performed at meta-
feature level (eg. gene level).
-F
(isGTFAnnotationFile)
Specify the format of the annotation file. Acceptable formats
include ‘GTF’ and ‘SAF’ (see Section 6.2.2 for details). By
default, C version of featureCounts program accepts a GTF
format annotation and R version accepts a SAF format anno-
tation. In-built annotations in SAF format are provided.
-g < string >
(GTF.attrType)
Specify the attribute type used to group features (eg. exons)
into meta-features (eg. genes) when GTF annotation is pro-
vided. ‘gene id’ by default. This attribute type is usually the
gene identifier. This argument is useful for the meta-feature
level summarization.
38
-G < string >
(genome)
Provide the name of a FASTA-format file that contains the
reference sequences used in read mapping that produced the
provided SAM/BAM files. This optional argument can be
used with ‘-J’ option to improve read counting for junctions.
-J
(juncCounts)
Count the number of reads supporting each exon-exon junc-
tion. Junctions will be identified from all the exon-spanning
reads (containing ‘N’ in CIGAR string) included in the input
data (note that options ‘–splitOnly’ and ‘–nonSplitOnly’ are
not considered by this parameter). The output result includes
names of primary and secondary genes that overlap at least
one of the two splice sites of a junction. Only one primary
gene is reported, but there might be more than one secondary
gene reported. Secondary genes do not overlap more splice
sites than the primary gene. When the primary and sec-
ondary genes overlap same number of splice sites, the gene
with the smallest leftmost base position is selected as the pri-
mary gene. Also included in the output result are the position
information for the left splice site (‘Site1’) and the right splice
site (‘Site2’) of a junction. These include chromosome name,
coordinate and strand of the splice site. In the last columns of
the output, number of supporting reads is provided for each
junction for each library.
-L
(isLongRead)
Turn on long-read counting mode. This option should be used
when counting long reads such as Nanopore or PacBio reads.
-M
(countMultiMappingReads)
If specified, multi-mapping reads/fragments will be counted.
The program uses the ‘NH’ tag to find multi-mapping reads.
Each alignment reported for a multi-mapping read will be
counted individually. Each alignment will carry 1 count or
a fractional count (--fraction). See section “Count multi-
mapping reads and multi-overlapping reads” for more details.
-o < string > Give the name of the output file. The output file contains
the number of reads assigned to each meta-feature (or each
feature if -f is specified). Note that the featureCounts function
in Rsubread does not use this parameter. It returns a list
object including read summarization results and other data.
39

-O
(allowMultiOverlap)
If specified, reads (or fragments) will be allowed to be assigned
to more than one matched meta-feature (or feature if -f is
specified). Reads/fragments overlapping with more than one
meta-feature/feature will be counted more than once. Note
that when performing meta-feature level summarization, a
read (or fragment) will still be counted once if it overlaps
with multiple features within the same meta-feature (as long
as it does not overlap with other meta-features). Also note
that this parameter is applied to each individual alignment
when there are more than one alignment reported for a read
(ie. multi-mapping read). See section “Count multi-mapping
reads and multi-overlapping reads” for more details.
-p
(isPairedEnd)
Specify that input data contain paired-end reads. feature-
Counts will terminate if the type of input reads (single-
end or paired-end) is different from the specified type. To
count fragments (instead of reads) for paired-end reads, the
--countReadPairs parameter should also be specified.
-P
(checkFragLength)
If specified, the fragment length will be checked when assign-
ing fragments to meta-features or features. This option is
only applicable for fragment counting. The fragment length
thresholds should be specified using -d and -D options.
-Q < int >
(minMQS)
The minimum mapping quality score a read must satisfy in
order to be counted. For paired-end reads, at least one end
should satisfy this criteria. 0 by default.
40

-R < string >
(reportReads)
Output detailed read assignment results for each read (or frag-
ment if paired end). The detailed assignment results can be
saved in three different formats including CORE, SAM and BAM
(note that these values are case sensitive).
When CORE format is specified, a tab-delimited file will be
generated for each input file. Name of each generated file is
the input file name added with ‘.featureCounts’. Each gen-
erated file contains four columns including read name, status
(assigned or the reason if not assigned), number of targets
and target list. A target is a feature or a meta-feature. Items
in the target lists is separated by comma. If a read is not
assigned, its number of targets will be set as -1.
When SAM or BAM format is specified, the detailed assignment
results will be saved to SAM and BAM format files. Names of
generated files are the input file names added with ‘.feature-
Counts.sam’ or ‘.featureCounts.bam’. Three tags are used to
describe read assignment results: XS, XN and XT. Tag XS
gives the assignment status. Tag XN gives number of targets.
Tag XT gives comma separated target list.
-s < intorstring >
(isStrandSpecific)
Indicate if strand-specific read counting should be performed.
A single integer value (applied to all input files) or a string
of comma-separated values (applied to each corresponding in-
put file) should be provided. Possible values include: 0 (un-
stranded), 1 (stranded) and 2 (reversely stranded). Default
value is 0 (ie. unstranded read counting carried out for all
input files). For paired-end reads, strand of the first read is
taken as the strand of the whole fragment. FLAG field is
used to tell if a read is first or second read in a pair. Value
of isStrandSpecific parameter in Rsubread featureCounts is
a vector which has a length of either 1, or the same with the
total number of input files provided.
-t < string >
(GTF.featureType)
Specify the feature type(s). If more than one feature type is
provided, they should be separated by ‘,’ (no space). Only
rows which have a matched feature type in the provided GTF
annotation file will be included for read counting. ‘exon’ by
default.
-T < int >
(nthreads)
Number of the threads. The value should be between 1 and
32. 1 by default.
-v Output version of the program.
41

−−byReadGroup
(byReadGroup)
Count reads by read group. Read group information is iden-
tified from the header of BAM/SAM input files and the gen-
erated count table will include counts for each group in each
library.
−−countReadPairs
(countReadPairs)
Read pairs will be counted instead of reads. This parameter
is only applicable when paired-end data were provided.
−−donotsort
(autosort)
If specified, paired end reads will not be re-ordered even if
reads from the same pair were found not to be next to each
other in the input.
−−extraAttributes
< string >
(GTF.attrType.extra)
Extract extra attribute types from the provided GTF annota-
tion and include them in the counting output. These attribute
types will not be used to group features. If more than one at-
tribute type is provided they should be separated by comma
(in Rsubread featureCounts its value is a character vector).
−−fraction
(fraction)
Assign fractional counts to features. This option must be used
together with ‘-M’ or ‘-O’ or both. When ‘-M’ is specified,
each reported alignment from a multi-mapping read (identi-
fied via ‘NH’ tag) will carry a count of 1/x, instead of 1 (one),
where x is the total number of alignments reported for the
same read. When ‘-O’ is specified, each overlapping feature
will receive a count of 1/y, where y is the total number of
features overlapping with the read. When both ‘-M’ and ‘-O’
are specified, each alignment will carry a count of 1/(x*y).
−−fracOverlap < float >
(fracOverlap)
Minimum fraction of overlapping bases in a read that is re-
quired for read assignment. Value should be a float number
in the range [0,1]. 0 by default. If paired end, number of over-
lapping bases is counted from both reads. Soft-clipped bases
are counted when calculating total read length (but ignored
when counting overlapping bases). Both this option and ‘–
minOverlap’ option need to be satisfied for read assignment.
−−fracOverlapFeature
< fl oat >
(fracOverlapFeature)
Minimum fraction of bases included in a feature that is re-
quired to overlap with a read or a read pair. Value should be
within range [0,1]. 0 by default.
−−ignoreDup
(ignoreDup)
If specified, reads that were marked as duplicates will be ig-
nored. Bit Ox400 in FLAG field of SAM/BAM file is used
for identifying duplicate reads. In paired end data, the entire
read pair will be ignored if at least one end is found to be a
duplicate read.
−−largestOverlap
(largestOverlap)
If specified, reads (or fragments) will be assigned to the target
that has the largest number of overlapping bases.
42

−−maxMOp < int >
(maxMOp)
Specify the maximum number of ‘M’ operations (matches or
mis-matches) allowed in a CIGAR string. 10 by default. Both
‘X’ and ‘=’ operations are treated as ‘M’ and adjacent ‘M’ op-
erations are merged in the CIGAR string. When the number
of ‘M’ operations exceeds the limit, only the first ‘maxMOp’
number of ‘M’ operations will be used in read assignment.
−−minOverlap < int >
(minOverlap)
Minimum number of overlapping bases in a read that is re-
quired for read assignment. 1 by default. If a negative value
is provided, then a gap of up to specified size will be allowed
between read and the feature that the read is assigned to. For
assignment of read pairs (fragments), number of overlapping
bases from each read from the same pair will be summed.
−−nonOverlap < int >
(nonOverlap)
Maximum number of non-overlapping bases in a read (or a
read pair) that is allowed when being assigned to a feature.
No limit is set by default.
−−nonOverlapFeature
< int >
(nonOverlapFeature)
Maximum number of non-overlapping bases in a feature that
is allowed in read assignment. No limit is set by default.
−−nonSplitOnly
(nonSplitOnly)
If specified, only non-split alignments (CIGAR strings do not
contain letter ‘N’) will be counted. All the other alignments
will be ignored.
−−primary
(primaryOnly)
If specified, only primary alignments will be counted. Primary
and secondary alignments are identified using bit 0x100 in
the Flag field of SAM/BAM files. All primary alignments
in a dataset will be counted no matter they are from multi-
mapping reads or not (ie. ‘-M’ is ignored).
−−read2pos < int >
(read2pos)
Read is reduced to its 5’ most base or 3’ most base. Read
summarization is then performed based on the single base
position to which the read is reduced. By default no read
reduction is performed. Read reduction is performed after
read shifting and read extension if they are also specified.
−−readExtension3 < int >
(readExtension3)
Reads are extended downstream by < int > bases from their
3’ end. 0 by default. Negative value is not allowed. Read
extension is performed after read shifting but before read re-
duction.
−−readExtension5 < int >
(readExtension5)
Reads are extended upstream by < int > bases from their 5’
end. 0 by default. Negative value is not allowed.
−−readShiftSize < int >
(readShiftSize)
Reads are shifted by < int > bases. 0 by default. Negative
value is not allowed.
43

−−readShiftType
< string >
(readShiftType)
Specify the direction in which reads are being shifted. Pos-
sible values include upstream, downstream, left and right.
upstream by default. Read shifting is performed before read
extension or reduction.
−−Rpath < string >
(reportReadsPath)
Specify a directory to save the detailed assignment results. If
unspecified, the directory where counting results are saved is
used. See ‘-R’ option for obtaining detailed assignment results
for reads.
−−splitOnly
(splitOnly)
If specified, only split alignments (CIGAR strings contain let-
ter ‘N’) will be counted. All the other alignments will be
ignored. An example of split alignments is the exon-spanning
reads in RNA-seq data. If exon-spanning reads need to be
assigned to all their overlapping exons, ‘-f’ and ‘-O’ options
should be provided as well.
−−tmpDir < string >
(tmpDir)
Directory under which intermediate files are saved (later re-
moved). By default, intermediate files will be saved to the
directory specified in ‘-o’ argument (In R, intermediate files
are saved to the current working directory by default).
−−verbose
(verbose)
Output verbose information for debugging such as unmatched
chromosomes/contigs between reads and annotation.
44

6.3 A quick start for featureCounts in SourceForge Sub-
read
You need to provide read mapping results (in either SAM or BAM format) and an annotation
file for the read summarization. The example commands below assume your annotation file
is in GTF format.
Summarize BAM format single-end reads using 5 threads:
featureCounts -T 5 -a annotation.gtf -t exon -g gene id
-o counts.txt mapping results SE.bam
Summarize BAM format single-end read data:
featureCounts -a annotation.gtf -t exon -g gene id
-o counts.txt mapping results SE.bam
Summarize multiple libraries at the same time:
featureCounts -a annotation.gtf -t exon -g gene id
-o counts.txt mapping results1.bam mapping results2.bam
Summarize paired-end reads and count fragments (instead of reads):
featureCounts -p --countReadPairs -a annotation.gtf -t exon -g gene id
-o counts.txt mapping results PE.bam
Count fragments satisfying the fragment length criteria, eg. [50bp, 600bp]:
featureCounts -p --countReadPairs -P -d 50 -D 600 -a annotation.gtf -t exon -g gene id
-o counts.txt mapping results PE.bam
Count fragments which have both ends successfully aligned without considering the fragment
length constraint:
featureCounts -p --countReadPairs -B -a annotation.gtf -t exon -g gene id
-o counts.txt mapping results PE.bam
Exclude chimeric fragments from the fragment counting:
featureCounts -p --countReadPairs -C -a annotation.gtf -t exon -g gene id
-o counts.txt mapping results PE.bam
45
6.4 A quick start for featureCounts in Bioconductor Rsub-
read
You need to provide read mapping results (in either SAM or BAM format) and an annotation
file for the read summarization. The example commands below assume your annotation file
is in GTF format.
Load Rsubread library from you R session:
library(Rsubread)
Summarize single-end reads using built-in RefSeq annotation for mouse genome ‘mm39’ (‘mm39’
is the default inbuilt genome annotation):
featureCounts(files="mapping_results_SE.bam")
Summarize single-end reads using a user-provided GTF annotation file:
featureCounts(files="mapping_results_SE.bam",annot.ext="annotation.gtf",
isGTFAnnotationFile=TRUE,GTF.featureType="exon",GTF.attrType="gene_id")
Summarize single-end reads using 5 threads:
featureCounts(files="mapping_results_SE.bam",nthreads=5)
Summarize BAM format single-end read data:
featureCounts(files="mapping_results_SE.bam")
Summarize multiple libraries at the same time:
featureCounts(files=c("mapping_results1.bam","mapping_results2.bam"))
Summarize paired-end reads and counting fragments (instead of reads):
featureCounts(files="mapping_results_PE.bam",isPairedEnd=TRUE)
Count fragments satisfying the fragment length criteria, eg. [50bp, 600bp]:
featureCounts(files="mapping_results_PE.bam",isPairedEnd=TRUE,checkFragLength=TRUE,
minFragLength=50,maxFragLength=600)
Count fragments which have both ends successfully aligned without considering the fragment
length constraint:
featureCounts(files="mapping_results_PE.bam",isPairedEnd=TRUE,requireBothEndsMapped=TRUE)
Exclude chimeric fragments from fragment counting:
featureCounts(files="mapping_results_PE.bam",isPairedEnd=TRUE,countChimericFragments=FALSE)
46

Chapter 7
Quantify 10x scRNA-seq data
The cellCounts program is developed for quantifying single-cell RNA-seq (scRNA-seq) data
generated by the 10x Genomics platform. With cellCounts, the entire quantification process
can be done by just one function call.
cellCounts takes raw scRNA-seq reads (BCL or FASTQ format) as input, maps them to the
reference genome and then produces UMI (Unique Molecular Identifier) counts for each gene in
each cell. The seed-and-vote paradigm is used in cellCounts read mapping. The featureCounts
function was adapted for read assignment performed within cellCounts. cellCounts also carries
out sample demultiplexing, read deduplication (UMI generation) and cell barcode calling
before generating a UMI count matrix. It can process multiple datasets at the same time.
Parameters of the cellCounts function are described below:
Table 4: Arguments used by the cellCounts program. The arguments are listed in the
alphabetical order.
Arguments Description
annot.ext Specify an external annotation for UMI counting. See
featureCounts function for more details. NULL by default.
annot.inbuilt Specify an inbuilt annotation for UMI counting. See
featureCounts function for more details. mm39 by default.
cell.barcode A character string giving the name of a text file (can be
gzipped) that contains the set of cell barcodes used in sample
preparation. If NULL, a cell barcode set will be determined for
the input data by cellCounts based on the matching of cell
barcodes sequences of the first 100,000 reads in the data with
the three cell barcode sets used by 10X Genomics. NULL by
default.
GTF.attrType See featureCounts function for more details. gene id by de-
fault.
GTF.featureType See featureCounts function for more details. exon by default.
index A character string giving the base name of index files gener-
ated for a reference genome by the buildindex function.
47

input.mode A character string specifying the input mode. The supported
input modes include BCL, FASTQ, FASTQ-dir and BAM. The BCL
mode includes BCL and CBCL formats, which are used by Illu-
mina for storing the raw reads directly generated from their
sequencers. When BCL mode is specified, cellCounts will au-
tomatically identify whether the input data are in BCL format
or CBCL format. FASTQ is the FASTQ format of sequencing
reads. FASTQ-dir is a directory where FASTQ-format reads
are saved. FASTQ-dir is useful for providing cellCounts the
FASTQ data generated by bcl2fastq program or bamtofastq
program (developed by 10X). BAM is the BAM format of
mapped read data with cell barcodes and UMI sequences in-
cluded. BCL by default.
isGTFAnnotationFile See featureCounts function for more details. FALSE by default.
maxMismatches Numeric value giving the maximum number of mismatched
bases allowed in the mapping of a read. 10 by default. Mis-
matches found in soft-clipped bases are not counted.
minMappedLength Numeric value giving the minimum number of mapped bases
in a read required for reporting a hit. 1 by default.
minVotes Numeric value giving the minimum number of votes required
for reporting a hit. 1 by default.
nBestLocations A numeric value giving the maximum number of reported
alignments for each multi-mapping read. 1 by default.
nsubreads Numeric value giving the number of subreads (seeds) ex-
tracted from each read. 15 by default.
nthreads A numeric value giving the number of threads used for read
mapping and counting. 10 by default.
reportExcludedBarcodes If TRUE, report UMI counts for those cell barcodes that were
filtered out during cell calling. FALSE by default.
48

sample A data frame or a character string providing sample-related
information. If the input format is BCL or CBCL, the provided
sample information should include the location where the
read data are stored, flowcell lanes used for sequencing, sam-
ple names and names of index sets used for indexing samples.
If a sample was sequenced in all lanes, then its lane number
can be set as *. Alternatively, all the lane numbers can be
listed for this sample. The sample information should be
saved to a data.frame object and then provided to the sample
parameter. Below shows an example of this data frame:
InputDirectory Lane SampleName IndexSetName
/path/to/dataset1 1 Sample1 SI-GA-E1
/path/to/dataset1 1 Sample2 SI-GA-E2
/path/to/dataset1 2 Sample1 SI-GA-E1
/path/to/dataset1 2 Sample2 SI-GA-E2
/path/to/dataset2 1 Sample3 SI-GA-E3
/path/to/dataset2 1 Sample4 SI-GA-E4
/path/to/dataset2 2 Sample3 SI-GA-E3
/path/to/dataset2 2 Sample4 SI-GA-E4
...
It is compulsory to have the four column headers shown
in the example above when generating this data frame for
a 10x dataset. If more than one datasets are provided for
analysis, the InputDirectory column should include more
than one distinct directory. Note that this data frame is
different from the Sample Sheet generated by the Illumina
sequencer. The cellCounts function uses the index set
names included in this data frame to generate an Illumina
Sample Sheet and then uses it to demultiplex all the samples.
If the input format is FASTQ, a data.frame object con-
taining the following three columns, BarcodeUMIFile,
ReadFile and SampleName, should be provided to the sample
parameter. Each row in the data frame represents a sample.
The BarcodeUMIFile column should include names of FASTQ
files containing cell barcode and UMI sequences for each read,
and the ReadFile column should include names of FASTQ
files containing genomic sequences for corresponding reads.
49

sample (cont’d) If the input format is FASTQ-dir, a character string, which
includes the path to the directory where the FASTQ-format
read data are stored, should be provided to the sample
parameter. The data in this directory are expected to be
generated by the bcl2fastq program (developed by Illu-
mina), or by the cellranger mkfastq command (developed
by 10x), or by the bamtofastq program (developed by 10x).
Finally, if the input format is BAM, a data.frame ob-
ject containing the following two columns, BAMFile and
SampleName, should be provided to the sample parameter.
Each row in the data frame represents a sample. The BAMFile
column should include names of BAM files containing
genomic sequences of reads with associated cell barcode and
UMI sequences. The cell barcode and UMI sequences should
be provided in the ‘CR’ and ‘UR’ tags included in the BAM
files. The Phred-encoded quality strings of the cell barcode
and UMI sequences should be provided in the ‘CY’ and ‘UY’
tags. These tags were originally defined in CellRanger BAM
files.
umi.cutoff Specify a UMI count cutoff for cell calling. All the cells with a
total UMI count greater than this cutoff will be called. If NULL,
a bootstrapping procedure will be performed to determine this
cutoff. NULL by default.
uniqueMapping A logical value indicating if only uniquely mapped reads
should be reported. FALSE by default.
useMetaFeatures Specify if UMI counting should be carried out at the meta-
feature level (eg. gene level). See featureCounts function for
more details. TRUE by default.
The cellCounts function returns a List object to R. It also outputs a BAM file for each sam-
ple. The BAM file includes location-sorted read mapping results. If the input mode is BCL or
BAM, it will also output a gzipped FASTQ file including cell barcode and UMI sequences (R1),
a gzipped FASTQ file including sample index sequences (I1), a gzipped FASTQ file including
sequences of the second sample index if dual-indexed library is used (I2) and a gzipped FASTQ
file including genomic sequences of the reads (R2).
The returned List object contains the following components:
counts
A List object including UMI counts for each sample. Each component in this object is a
matrix that contains UMI counts for a sample. Rows in the matrix are genes and columns are
cells.
annotation
50
A data.frame object containing a gene annotation. This is the annotation that was used for
the assignment of UMIs to genes during quantification. Rows in the annotation are genes.
Columns of the annotation include GeneID, Chr, Start, End and Length.
sample.info
A data.frame object containing sample information and quantification statistics. It includes
the following columns: SampleName, InputDirectory (if the input format is BCL), TotalCells,
HighConfidenceCells (if umi.cutoff is NULL), RescuedCells (if umi.cutoff is NULL), TotalUMI,
MinUMI, MedianUMI, MaxUMI, MeanUMI, TotalReads, MappedReads and AssignedReads. Each row
in the data frame is a sample.
cell.confidence
A List object indicating if a cell is a high-confidence cell or a rescued cell (low confidence).
Each component in the object is a logical vector indicating which cells in a sample are high-
confidence cells. cell.confidence is included in the output only if umi.cutoff is NULL.
counts.excluded.barcodes
A List object including UMI counts for excluded cell barcodes for each sample. Each UMI
count matrix is stored as a sparseMatrix object here.
51

Chapter 8
SNP calling
8.1 Algorithm
SNPs(Single Nucleotide Polymorphisms) are the mutations of single nucleotides in the genome.
It has been reported that many diseases were initiated and/or driven by such mutations.
Therefore, successful detection of SNPs is very useful in designing better diagnosis and treat-
ments for a variety of diseases such as cancer. SNP detection also is an important subject of
many population studies.
Next-gen sequencing technologies provide an unprecedented opportunity to identify SNPs
at the highest resolution. However, it is extremely computing-intensive to analyze the data
generated from these technologies for the purpose of SNP discovery because of the sheer
volume of the data and the large number of chromosomal locations to be considered. To
discover SNPs, reads need to be mapped to the reference genome first and then all the read
data mapped to a particular site will be used for SNP calling for that site. Discovery of SNPs is
often confounded by many sources of errors. Mapping errors and sequencing errors are often
the major sources of errors causing incorrect SNP calling. Incorrect alignments of indels,
exon-exon junctions and structural variants in the reads can also result in wrong placement
of blocks of continuous read bases, likely giving rise to consecutive incorrectly reported SNPs.
We have developed a highly accurate and efficient SNP caller, called exactSNP [10]. ex-
actSNP calls SNPs for individual samples, without requiring control samples to be provided.
It tests the statistical significance of SNPs by comparing SNP signals to their background
noises. It has been found to be an order of magnitude faster than existing SNP callers.
8.2 exactSNP
Below is the command for running exactSNP program. The complete list of parameters used
by exactSNP can be found in Table 5.
exactSNP [options] -i input -g reference genome -o output
52

Table 5: Arguments used by the exactSNP program included in the SourceForge Subread
package in alphabetical order. Arguments included in parenthesis are the equivalent
parameters used by exactSNP function in Bioconductor Rsubread package.
Arguments Description
-a < file >
(SNPAnnotationFile)
Specify name of a VCF-format file that includes annotated
SNPs. Such annotation files can be downloaded from public
databases such as the dbSNP database. Gzipped file is ac-
cepted. Incorporating known SNPs into SNP calling has been
found to be helpful. However note that the annotated SNPs
may or may not be called for the sample being analyzed.
-b
(isBAM)
Indicate the input file provided via −i is in BAM format.
-f < fl o at >
(minAllelicFraction)
Specify the minimum fraction of mis-matched bases a SNP-
containing location must have. Its value must between 0 and
1. 0 by default.
-g < file >
(refGenomeFile)
Specify name of the file including all reference sequences.
Only one single FASTA format file should be provided.
-i < file > [−b if BAM]
(readF ile)
Specify name of an input file including read mapping results.
The format of input file can be SAM or BAM (-b needs to be
specified if a BAM file is provided).
-n < int >
(minAllelicBases)
Specify the minimum number of mis-matched bases a SNP-
containing location must have. 1 by default.
-o < file >
(outputFile)
Specify name of the output file. This program outputs a VCF
format file that includes discovered SNPs.
-Q < int >
(qvalueCutoff)
Specify the q-value cutoff for SNP calling at sequencing depth
of 50X. 12 by default. The corresponding p-value cutoff is
10
−Q
. Note that this program automatically adjusts the q-
value cutoff according to the sequencing depth at each chro-
mosomal location.
-r < int >
(minReads)
Specify the minimum number of mapped reads a SNP-
containing location must have (ie. the minimum coverage).
1 by default.
-s < int >
(minBaseQuality)
Specify the cutoff for base calling quality scores (Phred scores)
read bases must satisfy to be used for SNP calling. 13 by
default. Read bases that have Phred scores lower than the
cutoff value will be excluded from the analysis.
-t < int >
(nTrimmedBases)
Specify the number of bases trimmed off from each end of the
read. 3 by default.
-T < int >
(nthreads)
Specify the number of threads. 1 by default.
-v Output version of the program.
53

-x < int >
(maxReads)
Specify the maximum depth a SNP location is allowed to have.
1,000,000 reads by default. Any location having more reads
than the maximum allowed depth will not be considered for
SNP calling. This option is useful for removing PCR artefacts.
54
Chapter 9
Utility programs
Usage info for each utility program can be seen by just typing the program name on the
command prompt.
9.1 repair
This program takes as input a paired-end BAM file and places reads from the same pair next
to each other in its output. BAM files generated by repair are compatible with featureCounts
program, ie they will not be re-sorted by featureCounts. Note that you do not have to run
repair before running featureCounts. featureCounts calls repair automatically if it finds that
reads need to be re-sorted.
The repair program uses a novel approach to quickly find reads from the same pair, rather
than performing time-consuming sort of read names. It takes only about half a minute to
re-order a location-sorted BAM file including 30 million read pairs.
9.2 flattenGTF
Flatten features (eg. exons) provided in a GTF annotation and output the modified annotation
to a SAF format annotation. If overlapping features are found in the GTF annotation, this
function can combine them to form a single large feature encompassing all the original features,
or chop them into non-overlapping bins.
9.3 promoterRegions
This function is only implemented in Rsubread. It generates a SAF format annotation that
includes coordinates of promoter regions for each gene.
55
9.4 propmapped
Get number of mapped reads from a BAM/SAM file.
9.5 qualityScores
Retrieve Phred scores for read bases from a Fastq/BAM/SAM file.
9.6 removeDup
Remove duplicated reads from a SAM/BAM file. In Rsubread this function is called re-
moveDupReads.
9.7 subread-fullscan
Get all chromosomal locations that contain a genomic sequence sharing high homology with
a given input sequence.
9.8 txUnique
This function is only implemented in Rsubread. It counts the number of bases unique to each
transcript.
56

Chapter 10
Case studies
10.1 A Bioconductor R pipeline for analyzing RNA-seq
data
Here we illustrate how to use two Bioconductor packages - Rsubread and limma - to perform a
complete RNA-seq analysis, including Subread read mapping, featureCounts read summariza-
tion, voom normalization and limma differential expresssion analysis.
Data and software. The RNA-seq data used in this case study include four libraries:
A 1, A 2, B 1 and B 2. Sample A is Universal Human Reference RNA (UHRR) and sample
B is Human Brain Reference RNA (HBRR). A 1 and A 2 are two replicates of sample A
(undergoing separate sample preparation), and B 1 and B 2 are two replicates of sample B.
In this case study, A 1 and A 2 are treated as biological replicates although they are more
like technical replicates. B 1 and B 2 are treated as biological replicates as well.
Note that these libraries only included reads originating from human chromosome 1 (ac-
cording to Subread aligner). Reads were generated by the MAQC/SEQC Consortium. Data
used in this case study can be downloaded by clicking here (283MB). Both read data and
reference sequence for chromosome 1 of human genome (GRCh37) were included in the data.
After downloading the data, you can uncompress it and save it to your current working
directory. Launch R and load Rsubread, limma and edgeR libraries by issuing the following
commands at your R prompt. Version of your R should be 3.0.2 or later. Rsubread version
should be 1.12.1 or later and limma version should be 3.18.0 or later. Note that this case study
only runs on Linux/Unix and Mac OS X.
library(Rsubread)
library(limma)
library(edgeR)
To install/update Rsubread and limma packages, issue the following commands at your R
prompt:
source("http://bioconductor.org/biocLite.R")
biocLite(pkgs=c("Rsubread","limma","edgeR"))
57
Index building. Build an index for human chromosome 1. This typically takes ∼3 minutes.
Index files with basename ‘chr1’ will be generated in your current working directory.
buildindex(basename="chr1",reference="hg19_chr1.fa")
Alignment. Perform read alignment for all four libraries and report uniquely mapped reads
only. This typically takes ∼5 minutes. BAM files containing the mapping results will be
generated in your current working directory.
targets <- readTargets()
align(index="chr1",readfile1=targets$InputFile,output_file=targets$OutputFile)
Read summarization. Summarize mapped reads to NCBI RefSeq genes. This will only
take a few seconds. Note that the featureCounts function contains built-in RefSeq annotations
for human and mouse genes. featureCounts returns an R ‘List’ object, which includes raw read
count for each gene in each library and also annotation information such as gene identifiers
and gene lengths.
fc <- featureCounts(files=targets$OutputFile,annot.inbuilt="hg19")
fc$counts[1:5,]
A_1.bam A_2.bam B_1.bam B_2.bam
653635 642 522 591 596
100422834 1 0 0 0
645520 5 3 0 0
79501 0 0 0 0
729737 82 72 30 25
fc$annotation[1:5,c("GeneID","Length")]
GeneID Length
1 653635 1769
2 100422834 138
3 645520 1130
4 79501 918
5 729737 3402
Create a DGEList object.
x <- DGEList(counts=fc$counts, genes=fc$annotation[,c("GeneID","Length")])
Filtering. Only keep in the analysis those genes which had >10 reads per million mapped
reads in at least two libraries.
isexpr <- rowSums(cpm(x) > 10) >= 2
x <- x[isexpr,]
Design matrix. Create a design matrix:
celltype <- factor(targets$CellType)
design <- model.matrix(~0+celltype)
colnames(design) <- levels(celltype)
58

Normalization. Perform voom normalization:
y <- voom(x,design,plot=TRUE)
The figure below shows the mean-variance relationship estimated by voom.
Sample clustering. Multi-dimensional scaling (MDS) plot shows that sample A libraries
are clearly separated from sample B libraries.
plotMDS(y,xlim=c(-2.5,2.5))
59

Linear model fitting and differential expression analysis. Fit linear models to genes
and assess differential expression using eBayes moderated t statistic. Here we compare sample
B vs sample A.
fit <- lmFit(y,design)
contr <- makeContrasts(BvsA=B-A,levels=design)
fit.contr <- eBayes(contrasts.fit(fit,contr))
dt <- decideTests(fit.contr)
summary(dt)
BvsA
-1 922
0 333
1 537
List top 10 differentially expressed genes:
options(digits=2)
topTable(fit.contr)
GeneID Length logFC AveExpr t P.Value adj.P.Val B
100131754 100131754 1019 1.6 16 113 3.5e-28 6.3e-25 54
2023 2023 1812 -2.7 13 -91 2.2e-26 1.9e-23 51
2752 2752 4950 2.4 13 82 1.5e-25 9.1e-23 49
22883 22883 5192 2.3 12 64 1.8e-23 7.9e-21 44
6135 6135 609 -2.2 12 -62 3.1e-23 9.5e-21 44
6202 6202 705 -2.4 12 -62 3.2e-23 9.5e-21 44
60
4904 4904 1546 -3.0 11 -60 5.5e-23 1.4e-20 43
23154 23154 3705 3.7 11 55 2.9e-22 6.6e-20 41
8682 8682 2469 2.6 12 49 2.2e-21 4.3e-19 39
6125 6125 1031 -2.0 12 -48 3.1e-21 5.6e-19 39
61
Bibliography
[1] Y. Liao, G. K. Smyth, and W. Shi. The subread aligner: fast, accurate and scalable read
mapping by seed-and-vote. Nucleic Acids Research, 41:e108, 2013.
[2] K. W. Tang, B. Alaei-Mahabadi, T. Samuelsson, M. Lindh, and E. Larsson. The land-
scape of viral expression and host gene fusion and adaptation in human cancer. Nature
Communications., 2013 Oct 1;4:2513. doi: 10.1038/ncomms3513, 2013.
[3] K. Man, M. Miasari, W. Shi, A. Xin, D. C. Henstridge, S. Preston, M. Pellegrini, G. T.
Belz, G. K. Smyth, M. A. Febbraio, S. L. Nutt, and A. Kallies. The transcription factor
IRF4 is essential for TCR affinity-mediated metabolic programming and clonal expansion
of T cells. Nature Immunology, 2013 Sep 22. doi: 10.1038/ni.2710, 2013.
[4] L. Spangenberg, P. Shigunov, A. P. Abud, A. R. Cofr´e, M. A. Stimamiglio, C. Kuligovski,
J. Zych, A. V. Schittini, A. D. Costa, C. K. Rebelatto, P. R. Brofman, S. Goldenberg,
A. Correa, H. Naya, and B. Dallagiovanna. Polysome profiling shows extensive posttran-
scriptional regulation during human adipocyte stem cell differentiation into adipocytes.
Stem Cell Research, 11:902–12, 2013.
[5] J. Z. Tang, C. L. Carmichael, W. Shi, D. Metcalf, A. P. Ng, C. D. Hyland, N. A. Jenkins,
N. G. Copeland, V. M. Howell, Z. J. Zhao, G. K. Smyth, B. T. Kile, and W. S. Alexander.
Transposon mutagenesis reveals cooperation of ETS family transcription factors with
signaling pathways in erythro-megakaryocytic leukemia. Proc Natl Acad Sci U S A,
110:6091–6, 2013.
[6] B. Pal, T. Bouras, W Shi, F. Vaillant, J. M. Sheridan, N. Fu, K. Breslin, K. Jiang, M. E.
Ritchie, M. Young, G. J. Lindeman, G. K. Smyth, and J. E. Visvader. Global changes in
the mammary epigenome are induced by hormonal cues and coordinated by Ezh2. Cell
Reports, 3:411–26, 2013.
[7] Y. Liao, G. K. Smyth, and W. Shi. featureCounts: an efficient general-purpose program
for assigning sequence reads to genomic features. Bioinformatics, 30:923–30, 2014.
[8] SEQC/MAQC-III Consortium. A comprehensive assessment of RNA-seq accuracy, re-
producibility and information content by the Sequencing Quality Control Consortium.
Nature Biotechnology, 32:903–14, 2014.
62
[9] Y. Liao, G. K. Smyth, and W. Shi. The R package Rsubread is easier, faster, cheaper
and better for alignment and quantification of RNA sequencing reads. Nucleic Acids
Research, 2019 Feb 20. doi: 10.1093/nar/gkz114. [Epub ahead of print], 2019.
[10] Y. Liao, G. K. Smyth, and W. Shi. ExactSNP: an efficient and accurate SNP calling
algorithm. In preparation.
63
